Alternative splicing affects the function and tissue-specific expression of the human constitutive androstane receptor
- PMID: 15043764
- PMCID: PMC406421
- DOI: 10.1186/1478-1336-2-1
Alternative splicing affects the function and tissue-specific expression of the human constitutive androstane receptor
Abstract
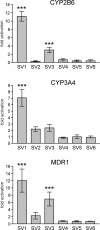
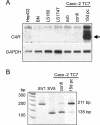
BACKGROUND: The constitutive androstane receptor (CAR) plays a key role in the control of drug metabolism and transport by mediating the phenobarbital-type induction of many phase I and II drug metabolizing enzymes and drug transporters. RESULTS: We identified transcripts generated by four different alternative splicing events in the human CAR gene. Two of the corresponding ligand binding domain isoforms demonstrated novel functional properties: First, CAR(SV3), which is encoded by a transcript containing an lengthened exon 7, differentially transactivated target gene promoters. Second, CAR(SV2), which results from the use of an alternative 3' splice site lengthening exon 8, showed ligand-dependent instead of constitutive interaction with coactivators. Furthermore, alternatively spliced transcripts demonstrated a tissue-specific expression pattern. In most tissues, only transcripts generated by alternative splicing within exon 9 were expressed. The encoded variant demonstrated a loss-of-function phenotype. Correct splicing of exon 8 to exon 9 is restricted to only a few tissues, among them liver and small intestine for which CAR function has been demonstrated, and is associated with the induction of CAR expression during differentiation of intestinal cells. CONCLUSION: Due to their specific activities, CAR variant proteins SV2 and SV3 may modulate the activity of reference CAR(SV1). Furthermore, we propose that transcriptional activation and regulation of splicing of exon 9 may be coupled to ensure appropriate tissue- and differentiation state-specific expression of transcripts encoding functional CAR protein. Altogether, alternative splicing seems to be of utmost importance for the regulation of CAR expression and function.
Figures





References
-
- Honkakoski P, Sueyoshi T, Negishi M. Drug-activated nuclear receptors CAR and PXR. Ann Med. 2003;35:172–182. - PubMed
LinkOut - more resources
Full Text Sources

