Successful herbivore attack due to metabolic diversion of a plant chemical defense
- PMID: 15051878
- PMCID: PMC387339
- DOI: 10.1073/pnas.0308007101
Successful herbivore attack due to metabolic diversion of a plant chemical defense
Abstract
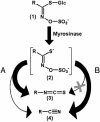
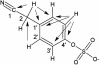
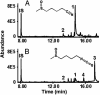
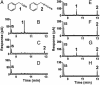
Plants protect themselves against herbivory with a diverse array of repellent or toxic secondary metabolites. However, many herbivorous insects have developed counteradaptations that enable them to feed on chemically defended plants without apparent negative effects. Here, we present evidence that larvae of the specialist insect, Pieris rapae (cabbage white butterfly, Lepidoptera: Pieridae), are biochemically adapted to the glucosinolate-myrosinase system, the major chemical defense of their host plants. The defensive function of the glucosinolate-myrosinase system results from the toxic isothiocyanates that are released when glucosinolates are hydrolyzed by myrosinases on tissue disruption. We show that the hydrolysis reaction is redirected toward the formation of nitriles instead of isothiocyanates if plant material is ingested by P. rapae larvae, and that the nitriles are excreted with the feces. The ability to form nitriles is due to a larval gut protein, designated nitrile-specifier protein, that by itself has no hydrolytic activity on glucosinolates and that is unrelated to any functionally characterized protein. Nitrile-specifier protein appears to be the key biochemical counteradaptation that allows P. rapae to feed with impunity on plants containing glucosinolates and myrosinases. This finding sheds light on the ecology and evolution of plant-insect interactions and suggests novel highly selective pest management strategies.
Figures


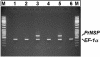
References
-
- Fahey, J. W., Zalcmann, A. T. & Talalay, P. (2001) Phytochemistry 56, 5–51. - PubMed
-
- Wittstock, U. & Halkier, B. A. (2002) Trends Plant Sci. 7, 263–270. - PubMed
-
- Rask, L., Andreasson, E., Ekbom, B., Eriksson, S., Pontoppidan, B. & Meijer, J. (2000) Plant Mol. Biol. 42, 93–113. - PubMed
-
- Benn, M. (1977) Pure Appl. Chem. 49, 197–210.
-
- Wittstock, U., Kliebenstein, D., Lambrix, V., Reichelt, M. & Gershenzon, J. (2003) Recent Adv. Phytochem. 37, 101–125.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

