DNA damage recognition of mutated forms of UvrB proteins in nucleotide excision repair
- PMID: 15065863
- PMCID: PMC1450103
- DOI: 10.1021/bi035992a
DNA damage recognition of mutated forms of UvrB proteins in nucleotide excision repair
Erratum in
- Biochemistry. 2004 Jun 1;43(21):6832
Abstract
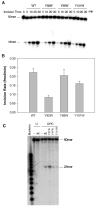
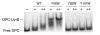
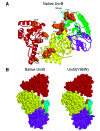
The DNA repair protein UvrB plays an indispensable role in the stepwise and sequential damage recognition of nucleotide excision repair in Escherichia coli. Our previous studies suggested that UvrB is responsible for the chemical damage recognition only upon a strand opening mediated by UvrA. Difficulties were encountered in studying the direct interaction of UvrB with adducts due to the presence of UvrA. We report herein that a single point mutation of Y95W in which a tyrosine is replaced by a tryptophan results in an UvrB mutant that is capable of efficiently binding to structure-specific DNA adducts even in the absence of UvrA. This mutant is fully functional in the UvrABC incisions. The dissociation constant for the mutant-DNA adduct interaction was less than 100 nM at physiological temperatures as determined by fluorescence spectroscopy. In contrast, similar substitutions at other residues in the beta-hairpin with tryptophan or phenylalanine do not confer UvrB such binding ability. Homology modeling of the structure of E. coli UvrB shows that the aromatic ring of residue Y95 and only Y95 directly points into the DNA binding cleft. We have also examined UvrB recognition of both "normal" bulky BPDE-DNA and protein-cross-linked DNA (DPC) adducts and the roles of aromatic residues of the beta-hairpin in the recognition of these lesions. A mutation of Y92W resulted in an obvious decrease in the efficiency of UvrABC incisions of normal adducts, while the incision of the DPC adduct is dramatically increased. Our results suggest that Y92 may function differently with these two types of adducts, while the Y95 residue plays an unique role in stabilizing the interaction of UvrB with DNA damage, most likely by a hydrophobic stacking.
Figures




References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

