Molecular epidemiology of Mycobacterium avium subsp. paratuberculosis isolates recovered from wild animal species
- PMID: 15071028
- PMCID: PMC387574
- DOI: 10.1128/JCM.42.4.1703-1712.2004
Molecular epidemiology of Mycobacterium avium subsp. paratuberculosis isolates recovered from wild animal species
Abstract
Mycobacterial isolates were obtained by radiometric culture from 33 different species of captive or free-ranging animals (n = 106) and environmental sources (n = 3) from six geographic zones within the United States. The identities of all 109 isolates were confirmed by using mycobactin J dependence and characterization of five well-defined molecular markers, including two integration loci of IS900 (loci L1 and L9), one Mycobacterium avium subsp. paratuberculosis (M. paratuberculosis)-specific sequence (locus 251), and one M. avium subsp. avium-specific marker (IS1245), as well as hsp65 and IS1311 restriction endonuclease analyses. Seventy-six acid-fast isolates were identified as M. paratuberculosis, 15 were identified as belonging to the M. avium-M. intracellulare complex (but not M. paratuberculosis), and the remaining 18 were identified as mycobacteria outside the M. avium-M. intracellulare complex. Fingerprinting by multiplex PCR for IS900 integration loci clustered 67 of the 76 M. paratuberculosis strains into a single clade (designated clade A18) and had a Simpson's diversity index (D) of 0.53. In contrast, sequence-based characterization of a recently identified M. paratuberculosis short sequence repeat (SSR) region enabled the differentiation of the M. paratuberculosis isolates in clade A18 into seven distinct alleles (D = 0.75). The analysis revealed eight subtypes among the 33 species of animals, suggesting the interspecies transmission of specific strains. Taken together, the results of our analyses demonstrate that SSR analysis enables the genetic characterization of M. paratuberculosis isolates from different host species and provide evidence for the host specificity of some M. paratuberculosis strains as well as sharing of strains between wild and domesticated animal species.
Figures
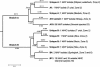

References
-
- Anonymous. 2001. CAST issue paper: Johne's disease in cattle. Council on Agriculture, Science and Technology Task Force. Council on Agriculture, Science and Technology, Washington, D.C.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

