A truncated aminoacyl-tRNA synthetase modifies RNA
- PMID: 15096612
- PMCID: PMC419641
- DOI: 10.1073/pnas.0401982101
A truncated aminoacyl-tRNA synthetase modifies RNA
Abstract
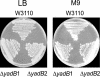
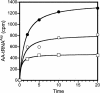
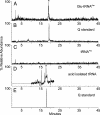
Aminoacyl-tRNA synthetases are modular enzymes composed of a central active site domain to which additional functional domains were appended in the course of evolution. Analysis of bacterial genome sequences revealed the presence of many shorter aminoacyl-tRNA synthetase paralogs. Here we report the characterization of a well conserved glutamyl-tRNA synthetase (GluRS) paralog (YadB in Escherichia coli) that is present in the genomes of >40 species of proteobacteria, cyanobacteria, and actinobacteria. The E. coli yadB gene encodes a truncated GluRS that lacks the C-terminal third of the protein and, consequently, the anticodon binding domain. Generation of a yadB disruption showed the gene to be dispensable for E. coli growth in rich and minimal media. Unlike GluRS, the YadB protein was able to activate glutamate in presence of ATP in a tRNA-independent fashion and to transfer glutamate onto tRNA(Asp). Neither tRNA(Glu) nor tRNA(Gln) were substrates. In contrast to canonical aminoacyl-tRNA, glutamate was not esterified to the 3'-terminal adenosine of tRNA(Asp). Instead, it was attached to the 2-amino-5-(4,5-dihydroxy-2-cyclopenten-1-yl) moiety of queuosine, the modified nucleoside occupying the first anticodon position of tRNA(Asp). Glutamyl-queuosine, like canonical Glu-tRNA, was hydrolyzed by mild alkaline treatment. Analysis of tRNA isolated under acidic conditions showed that this novel modification is present in normal E. coli tRNA; presumably it previously escaped detection as the standard conditions of tRNA isolation include an alkaline deacylation step that also causes hydrolysis of glutamyl-queuosine. Thus, this aminoacyl-tRNA synthetase fragment contributes to standard nucleotide modification of tRNA.
Figures



Comment in
-
Turning tRNA upside down: When aminoacylation is not a prerequisite to protein synthesis.Proc Natl Acad Sci U S A. 2004 May 18;101(20):7493-4. doi: 10.1073/pnas.0402276101. Epub 2004 May 11. Proc Natl Acad Sci U S A. 2004. PMID: 15138304 Free PMC article. No abstract available.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

