In planta engineering of viral RNA replicons: efficient assembly by recombination of DNA modules delivered by Agrobacterium
- PMID: 15103020
- PMCID: PMC406431
- DOI: 10.1073/pnas.0400149101
In planta engineering of viral RNA replicons: efficient assembly by recombination of DNA modules delivered by Agrobacterium
Erratum in
- Proc Natl Acad Sci U S A. 2004 Oct 26;101(43):15546
Abstract
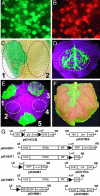
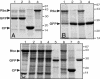
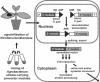
We have developed an efficient, versatile, and user-friendly viral engineering and expression system that is based on in planta assembly of functional viral vectors from separate pro-vector modules. With this new system, instead of supplying a plant cell with a complete viral vector as a mature viral particle, an RNA or a linear DNA molecule, we use agrobacteria to deliver various modules that are assembled inside the cell with the help of a site-specific recombinase. The resulting DNA is transcribed, and undesired elements such as recombination sites are spliced out, generating a fully functional RNA replicon. The proposed protocol allows us, by simply treating a plant with a mixture of two or more agrobacteria carrying specific prefabricated modules, to rapidly and inexpensively assemble and test multiple vector/gene combinations, without the need to perform the various engineering steps normally required with alternative protocols. The process described here is very fast (expression requires 3-4 days); it provides very high protein yield (up to 80% of total soluble protein); more than before, it is carried out using in vivo manipulations; it is based on prefabricated genetic modules that can be developed/upgraded independently; and it is inherently scalable.
Figures



References
-
- Yusibov, V., Shivprasad, S., Turpen, T. H., Dawson, W. & Koprowski, H. (1999) Curr. Top. Microbiol. Immunol. 240, 81-94. - PubMed
-
- Pogue, G. P., Lindbo, J. A., Garger, S. J. & Fitzmaurice, W. P. (2002) Annu. Rev. Phytopathol. 40, 45-74. - PubMed
-
- Shivprasad, S., Pogue, G. P., Lewandowski, D. J., Hidalgo, J., Donson, J., Grill, L. K. & Dawson, W. O. (1999) Virology 255, 312-323. - PubMed
-
- Fitzmaurice, W. P., Holzberg, S., Lindbo, J. A., Padgett, H. S., Palmer, K. E., Wolfe, G. M. & Pogue, G. P. (2002) OMICS 6, 137-151. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

