SIGI: score-based identification of genomic islands
- PMID: 15113412
- PMCID: PMC394314
- DOI: 10.1186/1471-2105-5-22
SIGI: score-based identification of genomic islands
Abstract
Background: Genomic islands can be observed in many microbial genomes. These stretches of DNA have a conspicuous composition with regard to sequence or encoded functions. Genomic islands are assumed to be frequently acquired via horizontal gene transfer. For the analysis of genome structure and the study of horizontal gene transfer, it is necessary to reliably identify and characterize these islands.
Results: A scoring scheme on codon frequencies Score_G1G2(cdn) = log(f_G2(cdn) / f_G1(cdn)) was utilized. To analyse genes of a species G1 and to test their relatedness to species G2, scores were determined by applying the formula to log-odds derived from mean codon frequencies of the two genomes. A non-redundant set of nearly 400 codon usage tables comprising microbial species was derived; its members were used alternatively at position G2. Genes having at least one score value above a species-specific and dynamically determined cut-off value were analysed further. By means of cluster analysis, genes were identified that comprise clusters of statistically significant size. These clusters were predicted as genomic islands. Finally and individually for each of these genes, the taxonomical relation among those species responsible for significant scores was interpreted. The validity of the approach and its limitations were made plausible by an extensive analysis of natural genes and synthetic ones aimed at modelling the process of gene amelioration.
Conclusions: The method reliably allows to identify genomic island and the likely origin of alien genes.
Figures
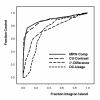
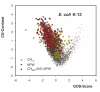

References
-
- Lobry JR. Asymmetric substitution patterns in the two DNA strands of bacteria. Mol Biol Evol. 1996;13:660–665. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources

