The splicing factor U2AF small subunit is functionally conserved between fission yeast and humans
- PMID: 15121844
- PMCID: PMC400479
- DOI: 10.1128/MCB.24.10.4229-4240.2004
The splicing factor U2AF small subunit is functionally conserved between fission yeast and humans
Abstract
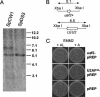
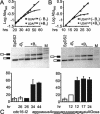
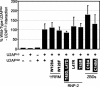
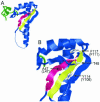
The small subunit of U2AF, which functions in 3' splice site recognition, is more highly conserved than its heterodimeric partner yet is less thoroughly investigated. Remarkably, we find that the small subunit of Schizosaccharomyces pombe U2AF (U2AF(SM)) can be replaced in vivo by its human counterpart, demonstrating that the conservation extends to function. Precursor mRNAs accumulate in S. pombe following U2AF(SM) depletion in a time frame consistent with a role in splicing. A comprehensive mutational analysis reveals that all three conserved domains are required for viability. Notably, however, a tryptophan in the pseudo-RNA recognition motif implicated in a key contact with the large subunit by crystallographic data is dispensable whereas amino acids implicated in RNA recognition are critical. Mutagenesis of the two zinc-binding domains demonstrates that they are neither equivalent nor redundant. Finally, two- and three-hybrid analyses indicate that mutations with effects on large-subunit interactions are rare whereas virtually all alleles tested diminished RNA binding by the heterodimer. In addition to demonstrating extraordinary conservation of U2AF small-subunit function, these results provide new insights into the roles of individual domains and residues.
Figures


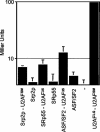
References
-
- Abovich, N., X. Liao, and M. Rosbash. 1994. The yeast MUD2 protein: an interaction with PRP11 defines a bridge between commitment complex and U2 snRNP addition. Genes Dev. 8:843-854. - PubMed
-
- Ambrozkova, M., F. Puta, L. Fukova, M. Skruzny, J. Brabek, and P. Folk. 2001. The fission yeast ortholog of the coregulator SKIP interacts with the small subunit of U2AF. Biochem. Biophys. Res. Commun. 284:1148-1154. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
