A novel family of P-loop NTPases with an unusual phyletic distribution and transmembrane segments inserted within the NTPase domain
- PMID: 15128444
- PMCID: PMC416466
- DOI: 10.1186/gb-2004-5-5-r30
A novel family of P-loop NTPases with an unusual phyletic distribution and transmembrane segments inserted within the NTPase domain
Abstract
Background: Recent sequence-structure studies on P-loop-fold NTPases have substantially advanced the existing understanding of their evolution and functional diversity. These studies provide a framework for characterization of novel lineages within this fold and prediction of their functional properties.
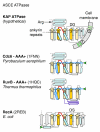
Results: Using sequence profile searches and homology-based structure prediction, we have identified a previously uncharacterized family of P-loop NTPases, which includes the neuronal membrane protein and receptor tyrosine kinase substrate Kidins220/ARMS, which is conserved in animals, the F-plasmid PifA protein involved in phage T7 exclusion, and several uncharacterized bacterial proteins. We refer to these (predicted) NTPases as the KAP family, after Kidins220/ARMS and PifA. The KAP family NTPases are sporadically distributed across a wide phylogenetic range in bacteria but among the eukaryotes are represented only in animals. Many of the prokaryotic KAP NTPases are encoded in plasmids and tend to undergo disruption to form pseudogenes. A unique feature of all eukaryotic and certain bacterial KAP NTPases is the presence of two or four transmembrane helices inserted into the P-loop NTPase domain. These transmembrane helices anchor KAP NTPases in the membrane such that the P-loop domain is located on the intracellular side. We show that the KAP family belongs to the same major division of the P-loop NTPase fold with the AAA+, ABC, RecA-like, VirD4-like, PilT-like, and AP/NACHT-like NTPase classes. In addition to the KAP family, we identified another small family of predicted bacterial NTPases, with two transmembrane helices inserted into the P-loop domain. This family is not specifically related to the KAP NTPases, suggesting independent acquisition of the transmembrane helices.
Conclusions: We predict that KAP family NTPases function principally in the NTP-dependent dynamics of protein complexes, especially those associated with the intracellular surface of cell membranes. Animal KAP NTPases, including Kidins220/ARMS, are likely to function as NTP-dependent regulators of the assembly of membrane-associated signaling complexes involved in neurite growth and development. One possible function of the prokaryotic KAP NTPases might be in the exclusion of selfish replicons, such as viruses, from the host cells. Phylogenetic analysis and phyletic patterns suggest that the common ancestor of the animals acquired a KAP NTPase via lateral transfer from bacteria. However, an earlier transfer into eukaryotes followed by multiple losses in several eukaryotic lineages cannot be ruled out.
Figures



References
-
- Koonin EV, Wolf YI, Aravind L. Protein fold recognition using sequence profiles and its application in structural genomics. Adv Protein Chem. 2000;54:245–275. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

