Seasonal changes in an alpine soil bacterial community in the colorado rocky mountains
- PMID: 15128545
- PMCID: PMC404381
- DOI: 10.1128/AEM.70.5.2867-2879.2004
Seasonal changes in an alpine soil bacterial community in the colorado rocky mountains
Abstract
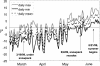
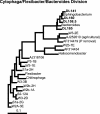
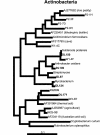
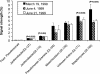
The period when the snowpack melts in late spring is a dynamic time for alpine ecosystems. The large winter microbial community begins to turn over rapidly, releasing nutrients to plants. Past studies have shown that the soil microbial community in alpine dry meadows of the Colorado Rocky Mountains changes in biomass, function, broad-level structure, and fungal diversity between winter and early summer. However, little specific information exists on the diversity of the alpine bacterial community or how it changes during this ecologically important period. We constructed clone libraries of 16S ribosomal DNA from alpine soil collected in winter, spring, and summer. We also cultivated bacteria from the alpine soil and measured the seasonal abundance of selected cultured isolates in hybridization experiments. The uncultured bacterial communities changed between seasons in diversity and abundance within taxa. The Acidobacterium division was most abundant in the spring. The winter community had the highest proportion of Actinobacteria and members of the Cytophaga/Flexibacter/Bacteroides (CFB) division. The summer community had the highest proportion of the Verrucomicrobium division and of beta-PROTEOBACTERIA: As a whole, alpha-Proteobacteria were equally abundant in all seasons, although seasonal changes may have occurred within this group. A number of sequences from currently uncultivated divisions were found, including two novel candidate divisions. The cultured isolates belonged to the alpha-, beta-, and gamma-Proteobacteria, the Actinobacteria, and the CFB groups. The only uncultured sequences that were closely related to the isolates were from winter and spring libraries. Hybridization experiments showed that actinobacterial and beta-proteobacterial isolates were most abundant during winter, while the alpha- and gamma-proteobacterial isolates tested did not vary significantly. While the cultures and clone libraries produced generally distinct groups of organisms, the two approaches gave consistent accounts of seasonal changes in microbial diversity.
Figures
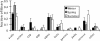





References
-
- Albrecht, W., A. Fischer, J. Smida, and E. Stackebrandt. 1987. Verrucomicrobium spinosum, a eubacterium representing an ancient line of descent. Syst. Appl. Microbiol. 10:57-62.
-
- Ausubel, F. M. (ed.). 1994. Current protocols in molecular biology. John Wiley and Sons, New York, N.Y.
-
- Bardgett, R. D., R. D. Lovell, P. J. Hobbs, and S. C. Jarvis. 1999. Seasonal changes in soil microbial communities along a fertility gradient of temperate grasslands. Soil Biol. Biochem. 31:1021-1030.
-
- Behrendt, U., A. Ulrich, P. Schumann, D. Naumann, and K. Suzuki. 2002. Diversity of grass-associated Microbacteriaceae isolated from the phyllosphere and litter layer after mulching the sward; polyphasic characterization of Subtercola pratensis sp. nov., Curtobacterium herbarum sp. nov. and Plantibacter flavus gen. nov., sp. nov. Int. J. Syst. Evol. Microbiol. 52:1441-1454. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

