Bifidobacterium lactis DSM 10140: identification of the atp (atpBEFHAGDC) operon and analysis of its genetic structure, characteristics, and phylogeny
- PMID: 15128574
- PMCID: PMC404453
- DOI: 10.1128/AEM.70.5.3110-3121.2004
Bifidobacterium lactis DSM 10140: identification of the atp (atpBEFHAGDC) operon and analysis of its genetic structure, characteristics, and phylogeny
Abstract
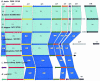
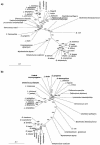
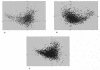
The atp operon is highly conserved among eubacteria, and it has been considered a molecular marker as an alternative to the 16S rRNA gene. PCR primers were designed from the consensus sequences of the atpD gene to amplify partial atpD sequences from 12 Bifidobacterium species and nine Lactobacillus species. All PCR products were sequenced and aligned with other atpD sequences retrieved from public databases. Genes encoding the subunits of the F(1)F(0)-ATPase of Bifidobacterium lactis DSM 10140 (atpBEFHAGDC) were cloned and sequenced. The deduced amino acid sequences of these subunits showed significant homology with the sequences of other organisms. We identified specific sequence signatures for the genus Bifidobacterium and for the closely related taxa Bifidobacterium lactis and Bifidobacterium animalis and Lactobacillus gasseri and Lactobacillus johnsonii, which could provide an alternative to current methods for identification of lactic acid bacterial species. Northern blot analysis showed that there was a transcript at approximately 7.3 kb, which corresponded to the size of the atp operon, and a transcript at 4.5 kb, which corresponded to the atpC, atpD, atpG, and atpA genes. The transcription initiation sites of these two mRNAs were mapped by primer extension, and the results revealed no consensus promoter sequences. Phylogenetic analysis of the atpD genes demonstrated that the Lactobacillus atpD gene clustered with the genera Listeria, Lactococcus, Streptococcus, and Enterococcus and that the higher G+C content and highly biased codon usage with respect to the genome average support the hypothesis that there was probably horizontal gene transfer. The acid inducibility of the atp operon of B. lactis DSM 10140 was verified by slot blot hybridization by using RNA isolated from acid-treated cultures of B. lactis DSM 10140. The rapid increase in the level of atp operon transcripts upon exposure to low pH suggested that the ATPase complex of B. lactis DSM 10140 was regulated at the level of transcription and not at the enzyme assembly step.
Figures


Similar articles
-
Analysis, characterization, and loci of the tuf genes in lactobacillus and bifidobacterium species and their direct application for species identification.Appl Environ Microbiol. 2003 Nov;69(11):6908-22. doi: 10.1128/AEM.69.11.6908-6922.2003. Appl Environ Microbiol. 2003. PMID: 14602655 Free PMC article.
-
Bifidobacterium lactis Meile et al. 1997 is a subjective synonym of Bifidobacterium animalis (Mitsuoka 1969) Scardovi and Trovatelli 1974.Microbiol Immunol. 2000;44(10):815-20. doi: 10.1111/j.1348-0421.2000.tb02568.x. Microbiol Immunol. 2000. PMID: 11128064
-
Biological and genetic classification of canine intestinal lactic acid bacteria and bifidobacteria.Microbiol Immunol. 2007;51(10):919-28. doi: 10.1111/j.1348-0421.2007.tb03983.x. Microbiol Immunol. 2007. PMID: 17951981
-
Comparative genomics of Bifidobacterium, Lactobacillus and related probiotic genera.Microb Ecol. 2012 Apr;63(3):651-73. doi: 10.1007/s00248-011-9948-y. Epub 2011 Oct 27. Microb Ecol. 2012. PMID: 22031452 Free PMC article. Review.
-
Taxonomy of Lactobacilli and Bifidobacteria.Curr Issues Intest Microbiol. 2007 Sep;8(2):44-61. Curr Issues Intest Microbiol. 2007. PMID: 17542335 Review.
Cited by
-
Functional genome analysis and anti-Helicobacter pylori activity of a novel bacteriocinogenic Lactococcus sp. NH2-7C from Thai fermented pork (Nham).Sci Rep. 2023 Nov 21;13(1):20362. doi: 10.1038/s41598-023-47687-4. Sci Rep. 2023. PMID: 37990119 Free PMC article.
-
Characterization of the groEL and groES loci in Bifidobacterium breve UCC 2003: genetic, transcriptional, and phylogenetic analyses.Appl Environ Microbiol. 2004 Oct;70(10):6197-209. doi: 10.1128/AEM.70.10.6197-6209.2004. Appl Environ Microbiol. 2004. PMID: 15466567 Free PMC article.
-
Microencapsulation in Alginate and Chitosan Microgels to Enhance Viability of Bifidobacterium longum for Oral Delivery.Front Microbiol. 2016 Apr 19;7:494. doi: 10.3389/fmicb.2016.00494. eCollection 2016. Front Microbiol. 2016. PMID: 27148184 Free PMC article.
-
Mechanism analysis of acid tolerance response of bifidobacterium longum subsp. longum BBMN 68 by gene expression profile using RNA-sequencing.PLoS One. 2012;7(12):e50777. doi: 10.1371/journal.pone.0050777. Epub 2012 Dec 7. PLoS One. 2012. PMID: 23236393 Free PMC article.
-
Gene structure and transcriptional organization of the dnaK operon of Bifidobacterium breve UCC 2003 and application of the operon in bifidobacterial tracing.Appl Environ Microbiol. 2005 Jan;71(1):487-500. doi: 10.1128/AEM.71.1.487-500.2005. Appl Environ Microbiol. 2005. PMID: 15640225 Free PMC article.
References
-
- Abrahams, J. P., A. G. W. Laslie, R. Lutter, and J. E. Walker. 1994. Structure at 2.8 A resolution of F1-ATPase from bovine heart mitochondria. Nature 370:621-628. - PubMed
-
- Arikado, E., H. Ishihara, T. Ehara, C. Shibata, H. Saito, T. Kakegawa, K. Igarashi, and H. Kobayashi. 1999. Enzyme level of enterococcal F1F0-ATPase is regulated by pH at the step of assembly. Eur. J. Biochem. 259:262-268. - PubMed
-
- Brochier, C., H. Philippe, and D. Moreira. 2000. The evolutionary history of ribosomal protein RpS14: horizontal gene transfer at the heart of the ribosome. Trends Genet. 16:529-533. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

