Highly significant linkage to the SLI1 locus in an expanded sample of individuals affected by specific language impairment
- PMID: 15133743
- PMCID: PMC1182086
- DOI: 10.1086/421529
Highly significant linkage to the SLI1 locus in an expanded sample of individuals affected by specific language impairment
Abstract
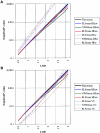
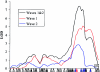
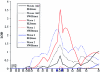
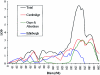
Specific language impairment (SLI) is defined as an unexplained failure to acquire normal language skills despite adequate intelligence and opportunity. We have reported elsewhere a full-genome scan in 98 nuclear families affected by this disorder, with the use of three quantitative traits of language ability (the expressive and receptive tests of the Clinical Evaluation of Language Fundamentals and a test of nonsense word repetition). This screen implicated two quantitative trait loci, one on chromosome 16q (SLI1) and a second on chromosome 19q (SLI2). However, a second independent genome screen performed by another group, with the use of parametric linkage analyses in extended pedigrees, found little evidence for the involvement of either of these regions in SLI. To investigate these loci further, we have collected a second sample, consisting of 86 families (367 individuals, 174 independent sib pairs), all with probands whose language skills are >/=1.5 SD below the mean for their age. Haseman-Elston linkage analysis resulted in a maximum LOD score (MLS) of 2.84 on chromosome 16 and an MLS of 2.31 on chromosome 19, both of which represent significant linkage at the 2% level. Amalgamation of the wave 2 sample with the cohort used for the genome screen generated a total of 184 families (840 individuals, 393 independent sib pairs). Analysis of linkage within this pooled group strengthened the evidence for linkage at SLI1 and yielded a highly significant LOD score (MLS = 7.46, interval empirical P<.0004). Furthermore, linkage at the same locus was also demonstrated to three reading-related measures (basic reading [MLS = 1.49], spelling [MLS = 2.67], and reading comprehension [MLS = 1.99] subtests of the Wechsler Objectives Reading Dimensions).
Figures
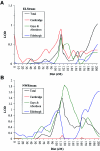
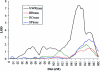
References
Electronic-Database Information
-
- Applied Biosystems, http://www.appliedbiosystems.com
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/
-
- UCSC, http://genome.ucsc.edu/
References
-
- Baddeley A, Wilson BA (1993) A developmental deficit in short-term phonological memory: implications for language and reading. Memory 1:65–78 - PubMed
-
- Baddeley AD, Gathercole SE, Papagno C (1998) The phonological loop as a language learning device. Psychol Rev 105:158–173 - PubMed
-
- Bartlett CW, Flax JF, Li W, Reaple-Bonilla T, Hayter J, Logue MW, Zimmerman R, Tallal P, Brzustowicz LM (2003) A genome scan of specific language impairment loci in families from the United States. Am J Hum Genet Suppl 73:A1884
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

