Functional analysis of the CXXC motif using phage antibodies that cross-react with protein disulphide-isomerase family proteins
- PMID: 15137910
- PMCID: PMC1133927
- DOI: 10.1042/BJ20040116
Functional analysis of the CXXC motif using phage antibodies that cross-react with protein disulphide-isomerase family proteins
Abstract
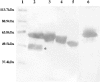
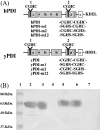
Polyclonal antibodies that had been raised against particular PDI (protein disulphide-isomerase) family proteins did not cross-react with other PDI family proteins. To evade immune tolerance to the important self-motif Cys-Xaa-Xaa-Cys, which is present in PDI family proteins, we used the phage display library [established by Griffiths, Williams, Hartley, Tomlinson, Waterhouse, Crosby, Kontermann, Jones, Low, Allison et al. (1994) EMBO J. 13, 3245-3260] to isolate successfully the phage antibodies that can cross-react with human and bovine PDIs, human P5, human PDI-related protein and yeast PDI. By measuring the binding of scFv (single-chain antibody fragment of variable region) to synthetic peptides and to mutants of PDI family proteins in a surface plasmon resonance apparatus, we identified clones that recognized sequences containing the CGHC motif or the CGHCK sequence. By using the isolated phage antibodies, we demonstrated for the first time that a lysine residue following the CXXC motif significantly increases the isomerase activities of PDI family proteins. Moreover, we demonstrated that the affinity of isolated scFvs for mutant PDI family proteins is proportional to the isomerase activities of their active sites.
Figures

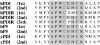

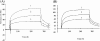


Similar articles
-
Evidence for mitochondrial localization of P5, a member of the protein disulphide isomerase family.J Biochem. 2008 Aug;144(2):187-96. doi: 10.1093/jb/mvn057. Epub 2008 Apr 19. J Biochem. 2008. PMID: 18424807
-
Anti-dengue virus nonstructural protein 1 antibodies recognize protein disulfide isomerase on platelets and inhibit platelet aggregation.Mol Immunol. 2009 Dec;47(2-3):398-406. doi: 10.1016/j.molimm.2009.08.033. Epub 2009 Oct 12. Mol Immunol. 2009. PMID: 19822367
-
The cell adhesion molecule retina cognin is a cell surface protein disulfide isomerase that uses disulfide exchange activity to modulate cell adhesion.Exp Cell Res. 2000 Jul 10;258(1):42-52. doi: 10.1006/excr.2000.4931. Exp Cell Res. 2000. PMID: 10912786
-
Two monoclonal antibodies to precisely the same epitope of type II collagen select non-crossreactive phage clones by phage display: implications for autoimmunity and molecular mimicry.Mol Immunol. 2004 Jun;41(4):411-9. doi: 10.1016/j.molimm.2004.03.025. Mol Immunol. 2004. PMID: 15163538
-
Potential roles of protein disulphide isomerase in viral infections.Acta Virol. 2013;57(3):293-304. Acta Virol. 2013. PMID: 24020755 Review.
Cited by
-
Protein disulfide isomerase in thrombosis and vascular inflammation.J Thromb Haemost. 2013 Dec;11(12):2084-91. doi: 10.1111/jth.12413. J Thromb Haemost. 2013. PMID: 24118938 Free PMC article. Review.
-
Identification and functional analysis of cadmium-binding protein in the visceral mass of Crassostrea gigas.Sci Rep. 2021 May 28;11(1):11306. doi: 10.1038/s41598-021-90882-4. Sci Rep. 2021. PMID: 34050239 Free PMC article.
-
Evolutionary Aspects of Selenium Binding Protein (SBP).J Mol Evol. 2023 Aug;91(4):471-481. doi: 10.1007/s00239-023-10105-4. Epub 2023 Apr 11. J Mol Evol. 2023. PMID: 37039856 Free PMC article.
-
Identification of novel autoantibodies for detection of malignant mesothelioma.PLoS One. 2013 Aug 19;8(8):e72458. doi: 10.1371/journal.pone.0072458. eCollection 2013. PLoS One. 2013. PMID: 23977302 Free PMC article.
-
Identification and enzymatic activities of four protein disulfide isomerase (PDI) isoforms of Leishmania amazonensis.Parasitol Res. 2008 Feb;102(3):437-46. doi: 10.1007/s00436-007-0784-4. Epub 2007 Dec 6. Parasitol Res. 2008. PMID: 18058133 Free PMC article.
References
-
- Freedman R. B., Hirst T. R., Tuite M. F. Protein disulphide isomerase: building bridges in protein folding. Trends Biochem. Sci. 1994;19:331–336. - PubMed
-
- Primm T. P., Walker K. W., Gilbert H. F. Facilitated protein aggregation. J. Biol. Chem. 1996;271:33664–33669. - PubMed
-
- Horibe T., Nagai H., Sakakibara K., Hagiwara Y., Kikuchi M. Ribostamycin inhibits the chaperone activity of protein disulfide isomerase. Biochem. Biophys. Res. Commun. 2001;289:967–972. - PubMed
-
- Horibe T., Nagai H., Matsui H., Hagiwara Y., Kikuchi M. Aminoglycoside antibiotics bind to protein disulfide isomerase and inhibit its chaperone activity. J. Antibiot. 2002;55:528–530. - PubMed
-
- Otsu M., Omura F., Yoshimori T., Kikuchi M. Protein disulfide isomerase associates with misfolded human lysozyme in vivo. J. Biol. Chem. 1994;269:6874–6877. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

