Metal-binding mechanism of Cox17, a copper chaperone for cytochrome c oxidase
- PMID: 15142040
- PMCID: PMC1133943
- DOI: 10.1042/BJ20040360
Metal-binding mechanism of Cox17, a copper chaperone for cytochrome c oxidase
Abstract
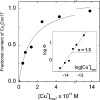
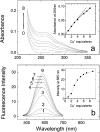
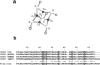
Cox17, a copper chaperone for cytochrome c oxidase, is an essential and highly conserved protein. The structure and mechanism of functioning of Cox17 are unknown, and even its metalbinding stoichiometry is elusive. In the present study, we demonstrate, using electrospray ionization-MS, that porcine Cox17 binds co-operatively four Cu+ ions. Cu4Cox17 is stable at pH values above 3 and fluorescence spectra indicate the presence of a solvent-shielded multinuclear Cu(I) cluster. Combining our results with earlier EXAFS results on yeast CuCox17, we suggest that Cu4Cox17 contains a Cu4S6-type cluster. At supramillimolar concentrations, dithiothreitol extracts metals from Cu4Cox17, and an apparent copper dissociation constant KCu=13 fM was calculated from these results. Charge-state distributions of different Cox17 forms suggest that binding of the first Cu+ ion to Cox17 causes a conformational change from an open to a compact state, which may be the rate-limiting step in the formation of Cu4Cox17. Cox17 binds non-co-operatively two Zn2+ ions, but does not bind Ag+ ions, which highlights its extremely high metal-binding specificity. We further demonstrate that porcine Cox17 can also exist in partly oxidized (two disulphide bridges) and fully oxidized (three disulphide bridges) forms. Partly oxidized Cox17 can bind one Cu+ or Zn2+ ion, whereas fully oxidized Cox17 does not bind metals. The metal-binding properties of Cox17 imply that, in contrast with other copper chaperones, Cox17 is designed for the simultaneous transfer of up to four copper ions to partner proteins. Metals can be released from Cox17 by non-oxidative as well as oxidative mechanisms.
Figures




Similar articles
-
Oxidative switches in functioning of mammalian copper chaperone Cox17.Biochem J. 2007 Nov 15;408(1):139-48. doi: 10.1042/BJ20070804. Biochem J. 2007. PMID: 17672825 Free PMC article.
-
Cox17, a copper chaperone for cytochrome c oxidase: expression, purification, and formation of mixed disulphide adducts with thiol reagents.Protein Expr Purif. 2007 May;53(1):138-44. doi: 10.1016/j.pep.2006.11.014. Epub 2006 Dec 5. Protein Expr Purif. 2007. PMID: 17208454
-
Characterization of the copper chaperone Cox17 of Saccharomyces cerevisiae.Biochemistry. 1998 May 19;37(20):7572-7. doi: 10.1021/bi980418y. Biochemistry. 1998. PMID: 9585572
-
Copper chaperones, intracellular copper trafficking proteins. Function, structure, and mechanism of action.Biochemistry (Mosc). 2003 Aug;68(8):827-37. doi: 10.1023/a:1025740228888. Biochemistry (Mosc). 2003. PMID: 12948382 Review.
-
Atx1-like chaperones and their cognate P-type ATPases: copper-binding and transfer.Biometals. 2007 Jun;20(3-4):275-89. doi: 10.1007/s10534-006-9068-1. Epub 2007 Jan 16. Biometals. 2007. PMID: 17225061 Review.
Cited by
-
Human Sco1 functional studies and pathological implications of the P174L mutant.Proc Natl Acad Sci U S A. 2007 Jan 2;104(1):15-20. doi: 10.1073/pnas.0606189103. Epub 2006 Dec 20. Proc Natl Acad Sci U S A. 2007. PMID: 17182746 Free PMC article.
-
Molecular mediators governing iron-copper interactions.Annu Rev Nutr. 2014;34:95-116. doi: 10.1146/annurev-nutr-071812-161215. Epub 2014 Jun 2. Annu Rev Nutr. 2014. PMID: 24995690 Free PMC article. Review.
-
Targeting the initiator to activate both ferroptosis and cuproptosis for breast cancer treatment: progress and possibility for clinical application.Front Pharmacol. 2025 Jan 10;15:1493188. doi: 10.3389/fphar.2024.1493188. eCollection 2024. Front Pharmacol. 2025. PMID: 39867656 Free PMC article. Review.
-
Function and redox state of mitochondrial localized cysteine-rich proteins important in the assembly of cytochrome c oxidase.Biochim Biophys Acta. 2008 Apr;1783(4):618-28. doi: 10.1016/j.bbamcr.2007.10.016. Epub 2007 Nov 9. Biochim Biophys Acta. 2008. PMID: 18070608 Free PMC article. Review.
-
Functional role of two interhelical disulfide bonds in human Cox17 protein from a structural perspective.J Biol Chem. 2011 Sep 30;286(39):34382-90. doi: 10.1074/jbc.M111.246223. Epub 2011 Aug 4. J Biol Chem. 2011. PMID: 21816817 Free PMC article.
References
-
- Ferguson-Miller S., Babcock G. T. Heme/copper terminal oxidases. Chem. Rev. 1996;96:2889–2908. - PubMed
-
- Tsukihara T., Aoyama H., Yamashita E., Tomizaki T., Yamaguchi H., Shinzawa-Itoh K., Nakashima R., Yaono R., Yoshikawa S. The whole structure of the 13-subunit oxidized cytochrome c oxidase at 2.8 Å. Science. 1996;272:1136–1144. - PubMed
-
- Huffman D. L., O'Halloran T. V. Function, structure, and mechanism of intracellular copper trafficking proteins. Annu. Rev. Biochem. 2001;70:677–701. - PubMed
-
- Glerum D. M., Shtanko A., Tzagoloff A. Characterization of Cox17, a yeast gene involved in copper metabolism and assembly of cytochrome oxidase. J. Biol. Chem. 1996;271:14504–14509. - PubMed
-
- Amaravadi R., Glerum D. M., Tzagoloff A. Isolation of a cDNA encoding the human homolog of Cox17, a yeast gene essential for mitochondrial copper recruitment. Hum. Genet. 1997;99:329–333. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

