The nuclear pore complex and the DEAD box protein Rat8p/Dbp5p have nonessential features which appear to facilitate mRNA export following heat shock
- PMID: 15143180
- PMCID: PMC416430
- DOI: 10.1128/MCB.24.11.4869-4879.2004
The nuclear pore complex and the DEAD box protein Rat8p/Dbp5p have nonessential features which appear to facilitate mRNA export following heat shock
Abstract
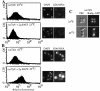
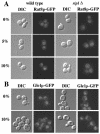
Nuclear pore complexes (NPCs) play an essential role in RNA export. Nucleoporins required for mRNA export in Saccharomyces cerevisiae are found in the Nup84p and Nup82p subcomplexes of the NPC. The Nup82p subcomplex contains Nup82p, Rat7p/Nup159p, Nsp1p, Gle1p/Rss1p, and Rip1p/Nup42p and is found only on the cytoplasmic face of NPCs. Both Rat7p and Gle1p contain binding sites for Rat8p/Dbp5p, an essential DEAD box protein and putative RNA helicase. Rip1p interacts directly with Gle1p and is the only protein known to be essential for mRNA export after heat shock but not under normal growth conditions. We report that in cells lacking Rip1p, both Gle1p and Rat8p dissociate from NPCs following heat shock at 42 degrees C. Rat8p but not Gle1p was retained at NPCs if rip1Delta cells were first shifted to 37 degrees C and then to 42 degrees C, and this was correlated with preserving mRNA export in heat-shocked rip1Delta cells. Export following ethanol shock was less dependent on the presence of Rip1p. Exposure to 10% ethanol led to dissociation of Rat8p from NPCs in both wild-type and rip1Delta cells. Following this treatment, Rat8p was primarily nuclear in wild-type cells but primarily cytoplasmic in rip1Delta cells. We also determined that efficient export of heat shock mRNA after heat shock depends upon a novel 6-amino-acid element within Rat8p. This motif is not required under normal growth conditions or following ethanol shock. These studies suggest that the molecular mechanism responsible for the defect in export of heat shock mRNAs in heat-shocked rip1Delta cells is dissociation of Rat8p from NPCs. These studies also suggest that both nuclear pores and Rat8p have features not required for mRNA export in growing cells but which enhance the ability of mRNAs to be exported following heat shock.
Figures





Similar articles
-
The RNA export factor Gle1p is located on the cytoplasmic fibrils of the NPC and physically interacts with the FG-nucleoporin Rip1p, the DEAD-box protein Rat8p/Dbp5p and a new protein Ymr 255p.EMBO J. 1999 Oct 15;18(20):5761-77. doi: 10.1093/emboj/18.20.5761. EMBO J. 1999. PMID: 10610322 Free PMC article.
-
Rat8p/Dbp5p is a shuttling transport factor that interacts with Rat7p/Nup159p and Gle1p and suppresses the mRNA export defect of xpo1-1 cells.EMBO J. 1999 Oct 15;18(20):5778-88. doi: 10.1093/emboj/18.20.5778. EMBO J. 1999. PMID: 10523319 Free PMC article.
-
Stress response in yeast mRNA export factor: reversible changes in Rat8p localization are caused by ethanol stress but not heat shock.J Cell Sci. 2004 Aug 15;117(Pt 18):4189-97. doi: 10.1242/jcs.01296. Epub 2004 Jul 27. J Cell Sci. 2004. PMID: 15280434
-
Nuclear export of messenger RNA.Results Probl Cell Differ. 2002;35:133-50. doi: 10.1007/978-3-540-44603-3_7. Results Probl Cell Differ. 2002. PMID: 11791404 Review. No abstract available.
-
Transport of messenger RNA from the nucleus to the cytoplasm.Curr Opin Cell Biol. 2006 Jun;18(3):299-306. doi: 10.1016/j.ceb.2006.04.006. Epub 2006 May 8. Curr Opin Cell Biol. 2006. PMID: 16682182 Review.
Cited by
-
Nup42 and IP6 coordinate Gle1 stimulation of Dbp5/DDX19B for mRNA export in yeast and human cells.Traffic. 2017 Dec;18(12):776-790. doi: 10.1111/tra.12526. Epub 2017 Oct 16. Traffic. 2017. PMID: 28869701 Free PMC article.
-
Gle1 is required for tRNA to stimulate Dbp5 ATPase activity in vitro and to promote Dbp5 mediated tRNA export in vivo.bioRxiv [Preprint]. 2023 Nov 9:2023.06.29.547072. doi: 10.1101/2023.06.29.547072. bioRxiv. 2023. Update in: Elife. 2024 Jan 08;12:RP89835. doi: 10.7554/eLife.89835. PMID: 37425677 Free PMC article. Updated. Preprint.
-
Following temperature stress, export of heat shock mRNA occurs efficiently in cells with mutations in genes normally important for mRNA export.Eukaryot Cell. 2007 Mar;6(3):505-13. doi: 10.1128/EC.00317-06. Epub 2007 Jan 26. Eukaryot Cell. 2007. PMID: 17259545 Free PMC article.
-
DDX55 promotes hepatocellular carcinoma progression by interacting with BRD4 and participating in exosome-mediated cell-cell communication.Cancer Sci. 2022 Sep;113(9):3002-3017. doi: 10.1111/cas.15393. Epub 2022 Jun 10. Cancer Sci. 2022. PMID: 35514200 Free PMC article.
-
RNA helicases and abiotic stress.Nucleic Acids Res. 2006 Jun 21;34(11):3220-30. doi: 10.1093/nar/gkl408. Print 2006. Nucleic Acids Res. 2006. PMID: 16790567 Free PMC article. Review.
References
-
- Bailer, S. M., C. Balduf, J. Katahira, A. Podtelejnikov, C. Rollenhagen, M. Mann, N. Pante, and E. Hurt. 2000. Nup116p associates with the Nup82p-Nsp1p-Nup159p nucleoporin complex. J. Biol. Chem. 275:23540-23548. - PubMed
-
- Cullen, B. R. 2003. Nuclear RNA export. J. Cell Sci. 116:587-597. - PubMed
-
- Damelin, M., and P. A. Silver. 2000. Mapping interactions between nuclear transport factors in living cells reveals pathways through the nuclear pore complex. Mol. Cell 5:133-140. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
