A novel partial modification at C2501 in Escherichia coli 23S ribosomal RNA
- PMID: 15146074
- PMCID: PMC1370582
- DOI: 10.1261/rna.5259404
A novel partial modification at C2501 in Escherichia coli 23S ribosomal RNA
Abstract
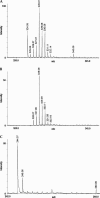
Escherichia coli is the best-characterized organism with respect to posttranscriptional modifications of its ribosomal RNA (rRNA). It is presently believed that all the modified nucleotides have been identified, primarily on the basis of two detection methods; modification-induced inhibition of the enzyme reverse transcriptase or analysis by combined HPLC and electrospray ionization mass spectrometry. Comparison of data from these different approaches reveals a disagreement regarding modification of C2501 in E. coli 23S rRNA. A. Bakin and J. Ofengand previously reported the detection of a modification at this site based on a reverse transcriptase assay. J.A. McCloskey and coworkers could not confirm the existence of such a modification using an electrospray ionization mass spectrometry approach. C2501 is therefore generally considered unmodified. We have used a strategy involving isolation of a specific rRNA fragment from E. coli 23S rRNA followed by Matrix Assisted Laser Desorption/Ionization mass spectrometry and tandem mass spectrometry to investigate this controversy. Our data reveal a novel 16-Da partial modification at C2501. We believe that the data reported here clarify the above discrepancy, because a minor partial modification detected in a reverse transcriptase assay would not necessarily be detected by the original mass spectrometry approach. The level of modification was furthermore monitored in different growth situations, and we found a significant positive regulation in stationary phase cells. C2501 is universally conserved and implicated in structure folds very close to the catalytic center of the ribosome. Moreover, several antibiotics bind to nucleotides in this region, which altogether make a modification at this site interesting.
Figures

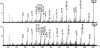

Similar articles
-
Posttranscriptional modifications in the A-loop of 23S rRNAs from selected archaea and eubacteria.RNA. 2002 Feb;8(2):202-13. doi: 10.1017/s1355838202013365. RNA. 2002. PMID: 11911366 Free PMC article.
-
Identification of 5-hydroxycytidine at position 2501 concludes characterization of modified nucleotides in E. coli 23S rRNA.J Mol Biol. 2011 Aug 19;411(3):529-36. doi: 10.1016/j.jmb.2011.06.036. Epub 2011 Jun 24. J Mol Biol. 2011. PMID: 21723290
-
UV-induced modifications in the peptidyl transferase loop of 23S rRNA dependent on binding of the streptogramin B antibiotic, pristinamycin IA.RNA. 1999 Apr;5(4):585-95. doi: 10.1017/s135583829998202x. RNA. 1999. PMID: 10199574 Free PMC article.
-
Mutational analysis of 23S ribosomal RNA structure and function in Escherichia coli.Adv Genet. 1999;41:157-95. doi: 10.1016/s0065-2660(08)60153-4. Adv Genet. 1999. PMID: 10494619 Review. No abstract available.
-
Comparison of functional peptide encoded in the Escherichia coli 23S rRNA with other peptides involved in cis-regulation of translation.Biochem Cell Biol. 1995 Nov-Dec;73(11-12):1061-70. doi: 10.1139/o95-114. Biochem Cell Biol. 1995. PMID: 8722022 Review.
Cited by
-
Identification of a novel methyltransferase, Bmt2, responsible for the N-1-methyl-adenosine base modification of 25S rRNA in Saccharomyces cerevisiae.Nucleic Acids Res. 2013 May 1;41(10):5428-43. doi: 10.1093/nar/gkt195. Epub 2013 Apr 4. Nucleic Acids Res. 2013. PMID: 23558746 Free PMC article.
-
Detection of internal N7-methylguanosine (m7G) RNA modifications by mutational profiling sequencing.Nucleic Acids Res. 2019 Nov 18;47(20):e126. doi: 10.1093/nar/gkz736. Nucleic Acids Res. 2019. PMID: 31504776 Free PMC article.
-
Biofilm Formation and Motility Are Promoted by Cj0588-Directed Methylation of rRNA in Campylobacter jejuni.Front Cell Infect Microbiol. 2018 Jan 18;7:533. doi: 10.3389/fcimb.2017.00533. eCollection 2017. Front Cell Infect Microbiol. 2018. PMID: 29404277 Free PMC article.
-
Aminoglycoside resistance 16S rRNA methyltransferases block endogenous methylation, affect translation efficiency and fitness of the host.RNA. 2014 Mar;20(3):382-91. doi: 10.1261/rna.042572.113. Epub 2014 Jan 7. RNA. 2014. PMID: 24398977 Free PMC article.
-
Bypassing rRNA methylation by RsmA/Dim1during ribosome maturation in the hyperthermophilic archaeon Nanoarchaeum equitans.Nucleic Acids Res. 2017 Feb 28;45(4):2007-2015. doi: 10.1093/nar/gkw839. Nucleic Acids Res. 2017. PMID: 28204608 Free PMC article.
References
-
- Bakin, A. and Ofengand, J. 1993. Four newly located pseudouridylate residues in Escherichia coli 23S ribosomal RNA are all at the peptidyltransferase center: Analysis by the application of a new sequencing technique. Biochemistry 32: 9754–9762. - PubMed
-
- Ban, N., Nissen, P., Hansen, J., Moore, P.B., and Steitz, T.A. 2000. The complete atomic structure of the large ribosomal subunit at 2.4 Å resolution. Science 289: 905–920. - PubMed
-
- Brimacombe, R., Mitchell, P., Osswald, M., Stade, K., and Bochkariov, D. 1993. Clustering of modified nucleotides at the functional center of bacterial ribosomal RNA. FASEB J. 7: 161–167. - PubMed
-
- Caldas, T., Binet, E., Bouloc, P., and Richarme, G. 2000. Translational defects of Escherichia coli mutants deficient in the Um(2552) 23S ribosomal RNA methyltransferase RrmJ/FTSJ. Biochem. Biophys. Res. Commun. 271: 714–718. - PubMed
-
- Cunningham, P.R., Richard, R.B., Weitzmann, C.J., Nurse, K., and Ofengand, J. 1991. The absence of modified nucleotides affects both in vitro assembly and in vitro function of the 30S ribosomal subunit of Escherichia coli. Biochimie 73: 789–796. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
