Multiple functions for the invariant AGC triad of U6 snRNA
- PMID: 15146076
- PMCID: PMC1370584
- DOI: 10.1261/rna.7310704
Multiple functions for the invariant AGC triad of U6 snRNA
Abstract
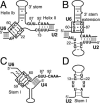
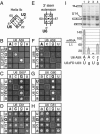
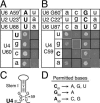
The invariant AGC triad of U6 snRNA plays an essential, unknown role in splicing. The triad has been implicated in base-pairing with residues in U2, U4, and U6. Through a genetic analysis in S. cerevisiae, we found that most AGC mutants are suppressed both by restoring pairing with U2, supporting the significance of U2/U6 helix Ib, and by destabilizing U2 stem I, indicating that this stem regulates helix Ib formation. Intriguingly, one of the helix Ib base pairs is required specifically for exon ligation, raising the possibility that the entirety of helix Ib is required only for exon ligation. We also found that U4 mutations that reduce complementarity in U4 stem I enhance U2-mediated suppression of an AGC mutant, suggesting that U4 stem I competes with the AGC-containing U4/U6 stem I. Implicating an additional, essential function for the triad, three triad mutants are refractory to suppression--even by simultaneous restoration of pairing with U2, U4, and U6. An absolute requirement for a purine at the central position of the triad parallels an equivalent requirement in a catalytically important AGC triad in group II introns, consistent with a role for the AGC triad of U6 in catalysis.
Figures

Similar articles
-
Spliceosome assembly in the absence of stable U4/U6 RNA pairing.RNA. 2015 May;21(5):923-34. doi: 10.1261/rna.048421.114. Epub 2015 Mar 11. RNA. 2015. PMID: 25762536 Free PMC article.
-
The conserved central domain of yeast U6 snRNA: importance of U2-U6 helix Ia in spliceosome assembly.RNA. 2002 Aug;8(8):997-1010. doi: 10.1017/s1355838202025013. RNA. 2002. PMID: 12212854 Free PMC article.
-
A stem/loop in U6 RNA defines a conformational switch required for pre-mRNA splicing.Genes Dev. 1994 Jan;8(2):221-33. doi: 10.1101/gad.8.2.221. Genes Dev. 1994. PMID: 8299941
-
A tertiary interaction detected in a human U2-U6 snRNA complex assembled in vitro resembles a genetically proven interaction in yeast.RNA. 2000 Feb;6(2):206-19. doi: 10.1017/s1355838200992197. RNA. 2000. PMID: 10688360 Free PMC article.
-
U2-U6 RNA folding reveals a group II intron-like domain and a four-helix junction.Nat Struct Mol Biol. 2004 Dec;11(12):1237-42. doi: 10.1038/nsmb863. Epub 2004 Nov 14. Nat Struct Mol Biol. 2004. PMID: 15543154
Cited by
-
Conformational heterogeneity of the protein-free human spliceosomal U2-U6 snRNA complex.RNA. 2013 Apr;19(4):561-73. doi: 10.1261/rna.038265.113. Epub 2013 Feb 20. RNA. 2013. PMID: 23426875 Free PMC article.
-
Emerging views about the molecular structure of the spliceosomal catalytic center.RNA Biol. 2012 Nov;9(11):1311-8. doi: 10.4161/rna.22359. Epub 2012 Oct 12. RNA Biol. 2012. PMID: 23064115 Free PMC article.
-
Evidence for a group II intron-like catalytic triplex in the spliceosome.Nat Struct Mol Biol. 2014 May;21(5):464-471. doi: 10.1038/nsmb.2815. Epub 2014 Apr 20. Nat Struct Mol Biol. 2014. PMID: 24747940 Free PMC article.
-
Termination of pre-mRNA splicing requires that the ATPase and RNA unwindase Prp43p acts on the catalytic snRNA U6.Genes Dev. 2019 Nov 1;33(21-22):1555-1574. doi: 10.1101/gad.328294.119. Epub 2019 Sep 26. Genes Dev. 2019. PMID: 31558568 Free PMC article.
-
U2 toggles iteratively between the stem IIa and stem IIc conformations to promote pre-mRNA splicing.Genes Dev. 2007 Apr 1;21(7):821-34. doi: 10.1101/gad.1536107. Genes Dev. 2007. PMID: 17403782 Free PMC article.
References
-
- Brow, D.A. and Guthrie, C. 1988. Spliceosomal RNA U6 is remarkably conserved from yeast to mammals. Nature 334: 213–218. - PubMed
-
- Fabrizio, P. and Abelson, J. 1990. Two domains of yeast U6 small nuclear RNA required for both steps of nuclear precursor messenger RNA splicing. Science 250: 404–409. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
