Xenopus U3 snoRNA docks on pre-rRNA through a novel base-pairing interaction
- PMID: 15146078
- PMCID: PMC1370586
- DOI: 10.1261/rna.5256704
Xenopus U3 snoRNA docks on pre-rRNA through a novel base-pairing interaction
Abstract
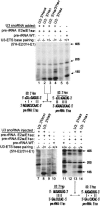
U3 small nucleolar RNA (snoRNA) is essential for rRNA processing to form 18S ribosomal RNA (rRNA). Previously, it has been shown that nucleolin is needed to load U3 snoRNA on pre-rRNA. However, as documented here, this is not sufficient. We present data that base-pairing between the U3 hinges and the external transcribed spacer (ETS) is critical for functional alignment of U3 on its pre-rRNA substrate. Additionally, the interaction between the U3 hinges and the ETS is proposed to serve as an anchor to hold U3 on the pre-rRNA substrate, while box A at the 5' end of U3 snoRNA swivels from ETS contacts to 18S rRNA contacts. Compensatory base changes revealed base-pairing between the 3' hinge of U3 snoRNA and region E1 of the ETS in Xenopus pre-rRNA; this novel interaction is required for 18S rRNA production. In contrast, base-pairing between the 5' hinge of U3 snoRNA and region E2 of the ETS is auxiliary, unlike the case in yeast where it is required. Thus, higher and lower eukaryotes use different interactions for functional association of U3 with pre-rRNA. The U3 hinge sequence varies between species, but covariation in the ETS retains complementarity. This species-specific U3-pre-rRNA interaction offers a potential target for a new class of antibiotics to prevent ribosome biogenesis in eukaryotic pathogens.
Figures




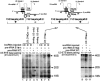


Similar articles
-
An evolutionary intra-molecular shift in the preferred U3 snoRNA binding site on pre-ribosomal RNA.Nucleic Acids Res. 2005 Sep 6;33(15):4995-5005. doi: 10.1093/nar/gki815. Print 2005. Nucleic Acids Res. 2005. PMID: 16147982 Free PMC article.
-
The spacing between functional Cis-elements of U3 snoRNA is critical for rRNA processing.J Mol Biol. 2000 Jun 30;300(1):57-74. doi: 10.1006/jmbi.2000.3798. J Mol Biol. 2000. PMID: 10864498
-
A second base pair interaction between U3 small nucleolar RNA and the 5'-ETS region is required for early cleavage of the yeast pre-ribosomal RNA.Nucleic Acids Res. 2011 Dec;39(22):9731-45. doi: 10.1093/nar/gkr675. Epub 2011 Sep 2. Nucleic Acids Res. 2011. PMID: 21890904 Free PMC article.
-
Comparative analysis of eukaryotic U3 snoRNA.RNA Biol. 2009 Nov-Dec;6(5):503-7. doi: 10.4161/rna.6.5.9607. RNA Biol. 2009. PMID: 19875933 Review.
-
Mpp10p, a new protein component of the U3 snoRNP required for processing of 18S rRNA precursors.Nucleic Acids Symp Ser. 1997;(36):64-7. Nucleic Acids Symp Ser. 1997. PMID: 9478208 Review.
Cited by
-
Analysis of sequence and structural features that identify the B/C motif of U3 small nucleolar RNA as the recognition site for the Snu13p-Rrp9p protein pair.Mol Cell Biol. 2007 Feb;27(4):1191-206. doi: 10.1128/MCB.01287-06. Epub 2006 Dec 4. Mol Cell Biol. 2007. PMID: 17145781 Free PMC article.
-
Functional regions in the 5' external transcribed spacer of yeast pre-rRNA.RNA. 2020 Jul;26(7):866-877. doi: 10.1261/rna.074807.120. Epub 2020 Mar 25. RNA. 2020. PMID: 32213618 Free PMC article.
-
Arabidopsis small nucleolar RNA monitors the efficient pre-rRNA processing during ribosome biogenesis.Proc Natl Acad Sci U S A. 2016 Oct 18;113(42):11967-11972. doi: 10.1073/pnas.1614852113. Epub 2016 Oct 5. Proc Natl Acad Sci U S A. 2016. PMID: 27708161 Free PMC article.
-
An evolutionary intra-molecular shift in the preferred U3 snoRNA binding site on pre-ribosomal RNA.Nucleic Acids Res. 2005 Sep 6;33(15):4995-5005. doi: 10.1093/nar/gki815. Print 2005. Nucleic Acids Res. 2005. PMID: 16147982 Free PMC article.
-
U3 snoRNA genes are multi-copy and frequently linked to U5 snRNA genes in Euglena gracilis.BMC Genomics. 2009 Nov 16;10:528. doi: 10.1186/1471-2164-10-528. BMC Genomics. 2009. PMID: 19917113 Free PMC article.
References
-
- Anderson, J.S., Lyon, C.E., Fox, A.H., Leung, A.K., Lam, Y.W., Steen, H., Mann, M., and Lamond, A.I. 2002. Directed proteomic analysis of the human nucleolus. Curr. Biol. 12: 1–11. - PubMed
-
- Borovjagin, A.V. and Gerbi, S.A. 1999. U3 small nucleolar RNA is essential for cleavage at sites 1, 2 and 3 in pre-rRNA and determines which rRNA processing pathway is taken in Xenopus oocytes. J. Mol. Biol. 286: 1347–1363. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
