A gene cluster for secondary metabolism in oat: implications for the evolution of metabolic diversity in plants
- PMID: 15148404
- PMCID: PMC419586
- DOI: 10.1073/pnas.0401301101
A gene cluster for secondary metabolism in oat: implications for the evolution of metabolic diversity in plants
Abstract
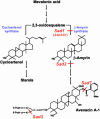
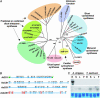
The evolution of the ability to synthesize specialized metabolites is likely to have been key for survival and diversification of different plant species. Oats (Avena spp.) produce antimicrobial triterpenoids (avenacins) that protect against disease. The oat beta-amyrin synthase gene AsbAS1, which encodes the first committed enzyme in the avenacin biosynthetic pathway, is clearly distinct from other plant beta-amyrin synthases. Here we show that AsbAS1 has arisen by duplication and divergence of a cycloartenol synthase-like gene, and that its properties have been refined since the divergence of oats and wheat. Strikingly, we have also found that AsbAS1 is clustered with other genes required for distinct steps in avenacin biosynthesis in a region of the genome that is not conserved in other cereals. Because the components of this gene cluster are required for at least four clearly distinct enzymatic processes (2,3-oxidosqualene cyclization, beta-amyrin oxidation, glycosylation, and acylation), it is unlikely that the cluster has arisen as a consequence of duplication of a common ancestor. Although clusters of paralogous genes are common in plants (e.g., gene clusters for rRNA and specific disease resistance), reports of clusters of genes that do not share sequence relatedness and whose products contribute to a single selectable function are rare [Gierl, A. & Frey, M. (2001) Planta 213, 493-498]. Taken together, our evidence has important implications for the generation of metabolic diversity in plants.
Figures

References
-
- Hostettmann, K. & Marston, A. (1995) Saponins (Cambridge Univ. Press, Cambridge, U.K.).
-
- Crombie, W. M. L. & Crombie, L. (1986) Phytochemistry 25, 2069–2073.
-
- Crombie, L., Crombie, W. M. L. & Whiting, D. A. (1986) J. Chem. Soc. Perkin Trans. 1, 1917–1922.
-
- Haralampidis, K., Trojanowska, M. & Osbourn, A. E. (2001) Adv. Biochem. Eng./Biotechnol. 75, 31–49. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

