Interactions between the nitrogen signal transduction protein PII and N-acetyl glutamate kinase in organisms that perform oxygenic photosynthesis
- PMID: 15150219
- PMCID: PMC415743
- DOI: 10.1128/JB.186.11.3346-3354.2004
Interactions between the nitrogen signal transduction protein PII and N-acetyl glutamate kinase in organisms that perform oxygenic photosynthesis
Erratum in
- J Bacteriol. 2004 Aug;186(16):5555
Abstract
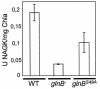
PII, one of the most conserved signal transduction proteins, is believed to be a key player in the coordination of nitrogen assimilation and carbon metabolism in bacteria, archaea, and plants. However, the identity of PII receptors remains elusive, particularly in photosynthetic organisms. Here we used yeast two-hybrid approaches to identify new PII receptors and to explore the extent of conservation of PII signaling mechanisms between eubacteria and photosynthetic eukaryotes. Screening of Synechococcus sp. strain PCC 7942 libraries with PII as bait resulted in identification of N-acetyl glutamate kinase (NAGK), a key enzyme in the biosynthesis of arginine. The integrity of Ser49, a residue conserved in PII proteins from organisms that perform oxygenic photosynthesis, appears to be essential for NAGK binding. The effect of glnB mutations on NAGK activity is consistent with positive regulation of NAGK by PII. Phylogenetic and yeast two-hybrid analyses strongly suggest that there was conservation of the NAGK-PII regulatory interaction in the evolution of cyanobacteria and chloroplasts, providing insight into the function of eukaryotic PII-like proteins.
Figures



References
-
- Atkinson, M. R., and A. J. Ninfa. 1999. Characterization of the GlnK protein of Escherichia coli. Mol. Microbiol. 32:301-313. - PubMed
-
- Ausubel, F. M. 1999. Short protocols in molecular biology: a compendium of methods from current protocols in molecular biology, 4th ed. Wiley, New York, N.Y.
-
- Bartel, P., C. T. Chien, R. Sternglanz, and S. Fields. 1993. Using the two-hybrid system to detect protein-protein interactions, p. 153-179. In D. A. Hartley (ed.), Cellular interactions in development: a practical approach. Oxford University Press, Oxford, United Kingdom.
-
- Blauwkamp, T. A., and A. J. Ninfa. 2003. Antagonism of PII signalling by the AmtB protein of Escherichia coli. Mol. Microbiol. 48:1017-1028. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

