Solution structure of the hypothetical protein Mth677 from Methanobacterium thermoautotrophicum: a novel alpha+beta fold
- PMID: 15152082
- PMCID: PMC2279996
- DOI: 10.1110/ps.04620504
Solution structure of the hypothetical protein Mth677 from Methanobacterium thermoautotrophicum: a novel alpha+beta fold
Abstract
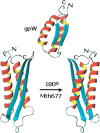
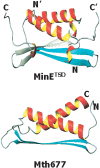
The structure of Mth677, a hypothetical protein from Methanobacterium thermoautotrophicum (Mth), has been determined by using heteronuclear nuclear magnetic resonance (NMR) methods on a double-labeled (15)N-(13)C sample. Mth677 adopts a novel alpha+beta fold, consisting of two alpha-helices (one N terminal and one C terminal) packed on the same side of a central beta-hairpin. This structure is likely shared by its three orthologs, detected in three other Archaebacteria. There are no clear features in the sequences of these proteins or in the genome organization of Mth to make a reliable functional assignment to this protein. However, the structural similarity to Escherichia coli MinE, the protein which controls that division occurs at the midcell site, lends support to the proposal that Mth677 might be, in Mth, the counterpart of the topological specificity domain of MinE in E. coli.
Figures



Similar articles
-
Structure of the archaeal translation initiation factor aIF2 beta from Methanobacterium thermoautotrophicum: implications for translation initiation.Protein Sci. 2004 Mar;13(3):659-67. doi: 10.1110/ps.03506604. Protein Sci. 2004. PMID: 14978306 Free PMC article.
-
Solution structure of the hypothetical protein MTH0637 from Methanobacterium thermoautotrophicum.J Biomol NMR. 2002 Mar;22(3):291-4. doi: 10.1023/a:1014908801766. J Biomol NMR. 2002. PMID: 11991357 No abstract available.
-
NMR structure determination and structure-based functional characterization of conserved hypothetical protein MTH1175 from Methanobacterium thermoautotrophicum.J Struct Funct Genomics. 2000;1(1):15-25. doi: 10.1023/a:1011348803324. J Struct Funct Genomics. 2000. PMID: 12836677
-
NMR-based structure of the conserved protein MTH865 from the archaeon Methanobacterium thermoautotrophicum.J Biomol NMR. 2001 Sep;21(1):63-6. doi: 10.1023/a:1011928105928. J Biomol NMR. 2001. PMID: 11693569 No abstract available.
-
Mass spectrometry of proteins of known mass.Proc Natl Acad Sci U S A. 2000 Dec 19;97(26):14025-7. doi: 10.1073/pnas.011526498. Proc Natl Acad Sci U S A. 2000. PMID: 11114170 Free PMC article. Review. No abstract available.
References
-
- Bax, A. and Grzesiek, S. 1993. Methodological advances in protein NMR. Acc. Chem. Res. 26 131–138.
-
- Bax, A., Clore, G.M., and Gronenborn, A.M. 1990. 1H-1H Correlation via isotropic mixing of 13C magnetization, a new three-dimensional approach for assigning 1H and 13C spectra of 13C enriched proteins. J. Mag. Reson. 88 425–431.
-
- Bax, A., Vuister, G.W., Grzesiek, S., Delaglio, F., Wang, A.C., Tschudin, R., and Zhu, G. 1994. Measurement of homo- and heteronuclear J couplings from quantitative J correlation. Methods Enzymol. 239 79–105. - PubMed
-
- Case, D.A., Pearlman, D.A., Caldwell, J.W., Cheatham III, T.E., Wang, J., Ross, W.S., Simmerling, C.L., Darden, T.A., Merz, K.M., et al. 2002. AMBER 7.0. University of California, San Francisco.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

