The sliding filament model: 1972-2004
- PMID: 15173218
- PMCID: PMC2234572
- DOI: 10.1085/jgp.200409089
The sliding filament model: 1972-2004
Figures


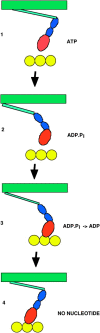
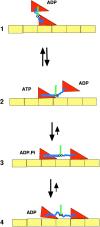
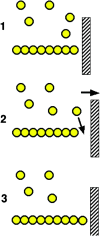

Similar articles
-
Mechanics and models of the myosin motor.Philos Trans R Soc Lond B Biol Sci. 2000 Apr 29;355(1396):433-40. doi: 10.1098/rstb.2000.0584. Philos Trans R Soc Lond B Biol Sci. 2000. PMID: 10836496 Free PMC article. Review.
-
The unit event of sliding of the chemo-mechanical enzyme composed of myosin and actin with regulatory proteins.Biochem Biophys Res Commun. 2008 Apr 25;369(1):144-8. doi: 10.1016/j.bbrc.2007.11.186. Epub 2007 Dec 26. Biochem Biophys Res Commun. 2008. PMID: 18157940 Review.
-
Lever arm model of force generation by actin-myosin-ATP.Biochemistry. 1999 Aug 3;38(31):9791-7. doi: 10.1021/bi9907633. Biochemistry. 1999. PMID: 10433684 Review. No abstract available.
-
Single-motor mechanics and models of the myosin motor.Philos Trans R Soc Lond B Biol Sci. 2000 Apr 29;355(1396):441-7. doi: 10.1098/rstb.2000.0585. Philos Trans R Soc Lond B Biol Sci. 2000. PMID: 10836497 Free PMC article. Review.
-
Stretching the lever-arm theory.Nature. 2002 Jan 10;415(6868):129-31. doi: 10.1038/415129a. Nature. 2002. PMID: 11805818 No abstract available.
Cited by
-
Dynamics of the nucleotide pocket of myosin measured by spin-labeled nucleotides.Biophys J. 2007 Jan 1;92(1):172-84. doi: 10.1529/biophysj.106.090035. Epub 2006 Oct 6. Biophys J. 2007. PMID: 17028139 Free PMC article.
-
Muscle contraction: A mechanical perspective.Eur Phys J E Soft Matter. 2010 Aug;32(4):411-8. doi: 10.1140/epje/i2010-10641-0. Epub 2010 Sep 7. Eur Phys J E Soft Matter. 2010. PMID: 20821341
-
The ATPase-dependent chaperoning activity of Hsp90a regulates thick filament formation and integration during skeletal muscle myofibrillogenesis.Development. 2008 Mar;135(6):1147-56. doi: 10.1242/dev.018150. Epub 2008 Feb 6. Development. 2008. PMID: 18256191 Free PMC article.
-
The interrelation between mechanical characteristics of contracting muscle, cross-bridge internal structure, and the mechanism of chemomechanical energy transduction.Eur Biophys J. 2012 Sep;41(9):733-53. doi: 10.1007/s00249-012-0849-x. Epub 2012 Aug 29. Eur Biophys J. 2012. PMID: 22930317
-
Post-translational modifications of vertebrate striated muscle myosin heavy chains.Cytoskeleton (Hoboken). 2024 Dec;81(12):832-842. doi: 10.1002/cm.21857. Epub 2024 Apr 8. Cytoskeleton (Hoboken). 2024. PMID: 38587113 Review.
References
-
- Bray, D. 1972. Cytoplasmic Actin: A Comparative Study. Cold Spring Harbor Symposia on Quantitative Biology. The Cold Spring Laboratory, Cold Spring Harbor. XXXVII:567–571.
-
- Case, R., D. Pierce, N. HomBooher, C. Hart, and R. Vale. 1997. The directional preference of kinesin motors is specified by an element outside of the motor catalytic domain. Cell. 90:959–966. - PubMed
-
- Cooke, R. 1986. The mechanism of muscle contraction. CRC Crit. Rev. Biochem. 21:53–118. - PubMed

