Heme regulates gene expression by triggering Crm1-dependent nuclear export of Bach1
- PMID: 15175654
- PMCID: PMC449764
- DOI: 10.1038/sj.emboj.7600248
Heme regulates gene expression by triggering Crm1-dependent nuclear export of Bach1
Erratum in
- EMBO J. 2014 Aug 1;33(15):1727
Abstract
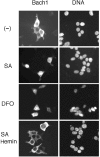
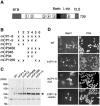
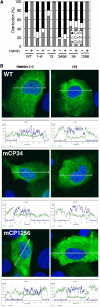
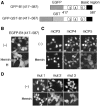
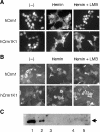
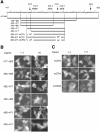
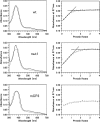
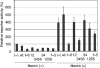
Bach1 is a transcriptional repressor of heme oxygenase-1 and beta-globin genes, both of which are known to be transcriptionally induced by heme. To test the hypothesis that heme regulates the activity of Bach1, we expressed wild type and mutated versions of Bach1 together with or without its heterodimer partner MafK in human 293T and GM02063 cells and examined their subcellular localization. Inhibition of heme synthesis enhanced the nuclear accumulation of Bach1, whereas treating cells with hemin resulted in nuclear exclusion of Bach1. While the cadmium-inducible nuclear export signal (NES) of Bach1 was dispensable for the heme response, a region containing two of the heme-binding motifs was found to be critical for the heme-induced nuclear exclusion. This region functioned as a heme-regulated NES dependent on the exporter Crm1. These results extend the regulatory roles for heme in protein sorting, and suggest that Bach1 transduces metabolic activity into gene expression.
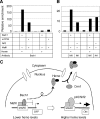
Figures

Similar articles
-
Heme-dependent up-regulation of the alpha-globin gene expression by transcriptional repressor Bach1 in erythroid cells.Biochem Biophys Res Commun. 2004 Nov 5;324(1):77-85. doi: 10.1016/j.bbrc.2004.09.022. Biochem Biophys Res Commun. 2004. PMID: 15464985
-
Heme mediates derepression of Maf recognition element through direct binding to transcription repressor Bach1.EMBO J. 2001 Jun 1;20(11):2835-43. doi: 10.1093/emboj/20.11.2835. EMBO J. 2001. PMID: 11387216 Free PMC article.
-
The role of Bach1 in ultraviolet A-mediated human heme oxygenase 1 regulation in human skin fibroblasts.Free Radic Biol Med. 2012 Jan 1;52(1):227-36. doi: 10.1016/j.freeradbiomed.2011.10.494. Epub 2011 Nov 6. Free Radic Biol Med. 2012. PMID: 22107958
-
Bach1, a heme-dependent transcription factor, reveals presence of multiple heme binding sites with distinct coordination structure.IUBMB Life. 2007 Aug-Sep;59(8-9):542-51. doi: 10.1080/15216540701225941. IUBMB Life. 2007. PMID: 17701549 Review.
-
Regulation of heme oxygenase-1 gene transcription: recent advances and highlights from the International Conference (Uppsala, 2003) on Heme Oxygenase.Antioxid Redox Signal. 2004 Oct;6(5):924-33. doi: 10.1089/ars.2004.6.924. Antioxid Redox Signal. 2004. PMID: 15345152 Review.
Cited by
-
Protective effect of tin chloride on rhabdomyolysis-induced acute kidney injury in rats.PLoS One. 2022 Mar 16;17(3):e0265512. doi: 10.1371/journal.pone.0265512. eCollection 2022. PLoS One. 2022. PMID: 35294485 Free PMC article.
-
The Heme Connection: Linking Erythrocytes and Macrophage Biology.Front Immunol. 2017 Jan 24;8:33. doi: 10.3389/fimmu.2017.00033. eCollection 2017. Front Immunol. 2017. PMID: 28167947 Free PMC article. Review.
-
Role of the tumor suppressor PTEN in antioxidant responsive element-mediated transcription and associated histone modifications.Mol Biol Cell. 2009 Mar;20(6):1606-17. doi: 10.1091/mbc.e08-07-0762. Epub 2009 Jan 21. Mol Biol Cell. 2009. PMID: 19158375 Free PMC article.
-
Analyzing the role of ferroptosis in ribosome-related bone marrow failure disorders: From pathophysiology to potential pharmacological exploitation.IUBMB Life. 2024 Dec;76(12):1011-1034. doi: 10.1002/iub.2897. Epub 2024 Jul 25. IUBMB Life. 2024. PMID: 39052023 Free PMC article. Review.
-
Antioxidant-induced phosphorylation of tyrosine 486 leads to rapid nuclear export of Bach1 that allows Nrf2 to bind to the antioxidant response element and activate defensive gene expression.J Biol Chem. 2010 Jan 1;285(1):153-62. doi: 10.1074/jbc.M109.040022. Epub 2009 Nov 6. J Biol Chem. 2010. PMID: 19897490 Free PMC article.
References
-
- Alam J, Shibahara S, Smith A (1989) Transcriptional activation of the heme oxygenase gene by heme and cadmium in mouse hepatoma cells. J Biol Chem 264: 6371–6375 - PubMed
-
- Alam J, Stewart D, Touchard C, Boinapally S, Choi AM, Cook JL (1999) Nrf2, a Cap‘n'Collar transcription factor, regulates induction of the heme oxygenase-1 gene. J Biol Chem 274: 26071–26078 - PubMed
-
- Alam J, Wicks C, Stewart D, Gong P, Touchard C, Otterbein S, Choi AMK, Burow ME, Tou J (2000) Mechanism of heme oxygenase-1 gene activation by cadmium in MCF-7 mammary epithelial cells. Role of p38 kinase and Nrf2 transcription factor. J Biol Chem 275: 27694–27702 - PubMed
-
- Andrews NC, Erdjument-Bromage H, Davidson MB, Tempst P, Orkin SH (1993a) Erythroid transcription factor NF-E2 is a haematopoietic-specific basic-leucine zipper protein. Nature 362: 722–728 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

