Proteomics for protein expression profiling in neuroscience
- PMID: 15176464
- PMCID: PMC3843356
- DOI: 10.1023/b:nere.0000023594.21352.17
Proteomics for protein expression profiling in neuroscience
Abstract
As the technology of proteomics moves from a theoretical approach to a practical reality, neuroscientists will have to determine the most appropriate applications for this technology. Neuroscientists will have to surmount difficulties particular to their research, such as limited sample amounts, heterogeneous cellular compositions in samples, and the fact that many proteins of interest are rare, hydrophobic proteins. This review examines protein isolation and protein fractionation and separation using two-dimensional electrophoresis (2-DE) and mass spectrometry proteomic methods. Methods for quantifying relative protein expression between samples (e.g., 2-DIGE, and ICAT) are also described. The coverage of the proteome, ability to detect membrane proteins, resource requirements, and quantitative reliability of different approaches is also discussed. Although there are many challenges in proteomic neuroscience, this field promises many rewards in the future.
Figures


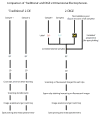
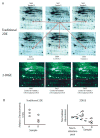

References
-
- Wilkins MR, Sanchez JC, Gooley AA, Appel RD, Humphery-Smith I, Hochstrasser DF, Williams KL. Progress with proteome projects: why all proteins expressed by a genome should be identified and how to do it. Biotechnol Genet Eng Rev. 1996;13:19–50. - PubMed
-
- Anderson L, Seilhamer J. A comparison of selected mRNA and protein abundances in human liver. Electrophoresis. 1997;18:533–537. - PubMed
-
- Mullis KB. Target amplification for DNA analysis by the polymerase chain reaction. Ann Biol Clin (Paris) 1990;48:579–582. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
