The circadian RNA-binding protein CHLAMY 1 represents a novel type heteromer of RNA recognition motif and lysine homology domain-containing subunits
- PMID: 15190002
- PMCID: PMC420122
- DOI: 10.1128/EC.3.3.815-825.2004
The circadian RNA-binding protein CHLAMY 1 represents a novel type heteromer of RNA recognition motif and lysine homology domain-containing subunits
Abstract
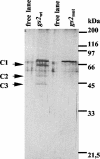
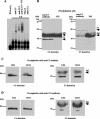
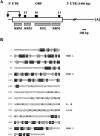
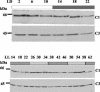
The RNA-binding protein CHLAMY 1 from Chlamydomonas reinhardtii binds specifically to UG> or =7 repeat sequences situated in the 3' untranslated regions of several mRNAs. Its binding activity is controlled by the circadian clock. The biochemical purification and characterization of CHLAMY 1 revealed a novel type of RNA-binding protein. It includes two different subunits (named C1 and C3), whose interaction appears necessary for RNA binding. One of them (C3) belongs to the proteins of the CELF (CUG-BP-ETR-3-like factors) family and thus bears three RNA recognition motif domains. The other is composed of three lysine homology domains and a protein-protein interaction domain (WW). The subunits C1 and C3 have theoretical molecular masses of 45 and 52 kDa, respectively, and are present in nearly equal amounts during the circadian cycle. At the beginning of the subjective night, both can be found in protein complexes of 100 to 160 kDa. However, during subjective day when binding activity of CHLAMY 1 is low, the C1 subunit in addition is present in a high-molecular-mass protein complex of more than 680 kDa. These data indicate posttranslational control of the circadian binding activity of CHLAMY 1. Notably, the C3 subunit shows significant homology to the rat CUG-binding protein 2. Anti-C3 antibodies can recognize the rat homologue, which can also be found in a protein complex in this vertebrate.
Copyright 2004 American Society for Microbiology
Figures
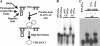




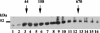
Similar articles
-
A heteromeric RNA-binding protein is involved in maintaining acrophase and period of the circadian clock.Plant Physiol. 2006 Oct;142(2):797-806. doi: 10.1104/pp.106.085944. Epub 2006 Aug 18. Plant Physiol. 2006. PMID: 16920878 Free PMC article.
-
Novel interaction of two clock-relevant RNA-binding proteins C3 and XRN1 in Chlamydomonas reinhardtii.FEBS Lett. 2012 Nov 16;586(22):3969-73. doi: 10.1016/j.febslet.2012.09.046. Epub 2012 Oct 12. FEBS Lett. 2012. PMID: 23068615
-
In vitro mutagenesis of binding site elements for the clock-controlled proteins CCTR and Chlamy 1.Biol Chem. 1997 Oct;378(10):1167-70. Biol Chem. 1997. PMID: 9372186
-
The function of circadian RNA-binding proteins and their cis-acting elements in microalgae.Chronobiol Int. 2003 Jul;20(4):529-41. doi: 10.1081/cbi-120022411. Chronobiol Int. 2003. PMID: 12916711 Review.
-
The circadian clock of the unicellular eukaryotic model organism Chlamydomonas reinhardtii.Biol Chem. 2003 May;384(5):689-95. doi: 10.1515/BC.2003.077. Biol Chem. 2003. PMID: 12817465 Review.
Cited by
-
The enigmatic clock of dinoflagellates, is it unique?Front Microbiol. 2022 Oct 19;13:1004074. doi: 10.3389/fmicb.2022.1004074. eCollection 2022. Front Microbiol. 2022. PMID: 36338102 Free PMC article. Review.
-
Evolution of circadian clocks along the green lineage.Plant Physiol. 2022 Sep 28;190(2):924-937. doi: 10.1093/plphys/kiac141. Plant Physiol. 2022. PMID: 35325228 Free PMC article.
-
Predicting the physiological role of circadian metabolic regulation in the green alga Chlamydomonas reinhardtii.PLoS One. 2011;6(8):e23026. doi: 10.1371/journal.pone.0023026. Epub 2011 Aug 22. PLoS One. 2011. PMID: 21887226 Free PMC article.
-
Analysis of flagellar phosphoproteins from Chlamydomonas reinhardtii.Eukaryot Cell. 2009 Jul;8(7):922-32. doi: 10.1128/EC.00067-09. Epub 2009 May 8. Eukaryot Cell. 2009. PMID: 19429781 Free PMC article.
-
Diversity of plant circadian clocks: Insights from studies of Chlamydomonas reinhardtii and Physcomitrella patens.Plant Signal Behav. 2016;11(1):e1116661. doi: 10.1080/15592324.2015.1116661. Plant Signal Behav. 2016. PMID: 26645746 Free PMC article.
References
-
- Adams, P. D., S. Seeholzer, and M. Ohn. 2002. Identification of associated proteins by coimmunoprecipitation. Protein-protein interaction: a molecular cloning manual, p. 59-74. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
-
- Adinolfi, S., A. Ramos, S. R. Martin, P. F. Dal, P. Pucci, B. Bardoni, J. L. Mandel, and A. Pastore. 2003. The N-terminus of the fragile X mental retardation protein contains a novel domain involved in dimerization and RNA binding. Biochemistry 42:10437-10444. - PubMed
-
- Allada, R., P. Emery, J. S. Takahashi, and M. Rosbash. 2001. Stopping time: the genetics of fly and mouse circadian clocks. Annu. Rev. Neurosci. 24:1091-1119. - PubMed
-
- Buratti, E., and F. B. Baralle. 2001. Characterization and functional implications of the RNA binding properties of nuclear factor TDP-43, a novel splicing regulator of CFTR exon 9. J. Biol. Chem. 276:36337-36343. - PubMed
-
- Burd, C. G., and G. Dreyfuss. 1994. Conserved structures and diversity of functions of RNA-binding proteins. Science 265:615-621. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

