Selection pressure-driven evolution of the Epstein-Barr virus-encoded oncogene LMP1 in virus isolates from Southeast Asia
- PMID: 15194789
- PMCID: PMC421669
- DOI: 10.1128/JVI.78.13.7131-7137.2004
Selection pressure-driven evolution of the Epstein-Barr virus-encoded oncogene LMP1 in virus isolates from Southeast Asia
Abstract
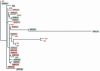
The geographically constrained distribution of Epstein-Barr virus (EBV)-associated nasopharyngeal carcinoma (NPC) in southeast Asian populations suggests that both viral and host genetics may influence disease risk. Although susceptibility loci have been mapped within the human genome, the role of viral genetics in the focal distribution of NPC remains an enigma. Here we report a molecular phylogenetic analysis of an NPC-associated viral oncogene, LMP1, in a large panel of EBV isolates from southeast Asia and from Papua New Guinea, Africa, and Australia, regions of the world where NPC is and is not endemic, respectively. This analysis revealed that LMP1 sequences show a distinct geographic structure, indicating that the southeast Asian isolates have evolved as a lineage distinct from those of Papua New Guinea, African, and Australian isolates. Furthermore, a likelihood ratio test revealed that the C termini of the LMP1 sequences of the southeast Asian lineage are under significant positive selection pressure, particularly at some sites within the C-terminal activator regions. We also present evidence that although the N terminus and transmembrane region of LMP1 have undergone recombination, the C-terminal region of the gene has evolved without any history of recombination. Based on these observations, we speculate that selection pressure may be driving the LMP1 sequences in virus isolates from southeast Asia towards a more malignant phenotype, thereby influencing the endemic distribution of NPC in this region.
Figures


Similar articles
-
Nucleotide sequences and functions of the Epstein-Barr virus latent membrane protein 1 genes isolated from salivary gland lymphoepithelial carcinomas.Virus Genes. 2005 Mar;30(2):223-35. doi: 10.1007/s11262-004-5630-5. Virus Genes. 2005. PMID: 15744579
-
Epstein-Barr virus genetic variation in Vietnamese patients with nasopharyngeal carcinoma: full-length analysis of LMP1.Virus Genes. 2008 Oct;37(2):273-81. doi: 10.1007/s11262-008-0262-9. Epub 2008 Jul 29. Virus Genes. 2008. PMID: 18663567
-
Modified Anoikis Assay That Functionally Segregates Epstein-Barr Virus LMP1 Strains into Two Groups.J Virol. 2018 Aug 29;92(18):e00557-18. doi: 10.1128/JVI.00557-18. Print 2018 Sep 15. J Virol. 2018. PMID: 29950426 Free PMC article.
-
Pathogenic role of Epstein-Barr virus latent membrane protein-1 in the development of nasopharyngeal carcinoma.Cancer Lett. 2013 Aug 28;337(1):1-7. doi: 10.1016/j.canlet.2013.05.018. Epub 2013 May 17. Cancer Lett. 2013. PMID: 23689138 Review.
-
Progression of understanding for the role of Epstein-Barr virus and management of nasopharyngeal carcinoma.Cancer Metastasis Rev. 2017 Sep;36(3):435-447. doi: 10.1007/s10555-017-9693-x. Cancer Metastasis Rev. 2017. PMID: 28819752 Free PMC article. Review.
Cited by
-
Nucleotide sequences and functions of the Epstein-Barr virus latent membrane protein 1 genes isolated from salivary gland lymphoepithelial carcinomas.Virus Genes. 2005 Mar;30(2):223-35. doi: 10.1007/s11262-004-5630-5. Virus Genes. 2005. PMID: 15744579
-
The vIL-10 gene of the Epstein-Barr virus (EBV) is conserved in a stable manner except for a few point mutations in various EBV isolates.Virus Genes. 2007 Dec;35(3):563-9. doi: 10.1007/s11262-007-0153-5. Epub 2007 Aug 31. Virus Genes. 2007. PMID: 17763933
-
Positive selection on a bacterial oncoprotein associated with gastric cancer.Gut Pathog. 2011 Nov 11;3:18. doi: 10.1186/1757-4749-3-18. Gut Pathog. 2011. PMID: 22078307 Free PMC article.
-
Epstein-Barr virus isolates retain their capacity to evade T cell immunity through BNLF2a despite extensive sequence variation.J Virol. 2012 Jan;86(1):572-7. doi: 10.1128/JVI.05151-11. Epub 2011 Oct 19. J Virol. 2012. PMID: 22013037 Free PMC article.
-
PD-L1 predicts poor prognosis for nasopharyngeal carcinoma irrespective of PD-1 and EBV-DNA load.Sci Rep. 2017 Mar 3;7:43627. doi: 10.1038/srep43627. Sci Rep. 2017. PMID: 28256540 Free PMC article.
References
-
- Awadalla, P., A. Eyre-Walker, and J. Maynard-Smith. 1999. Linkage disequilibrium and recombination in hominid mitochondrial DNA. Science 286:2524-2525. - PubMed
-
- Cheung, S. T., S. F. Leung, K. W. Lo, K. W. Chiu, J. S. Tam, T. F. Fok, P. J. Johnson, J. C. Lee, and D. P. Huang. 1998. Specific latent membrane protein 1 gene sequences in type 1 and type 2 Epstein-Barr virus from nasopharyngeal carcinoma in Hong Kong. Int. J. Cancer 76:399-406. - PubMed
-
- Edwards, R. H., F. Seillier-Moiseiwitsch, and N. Raab-Traub. 1999. Signature amino acid changes in latent membrane protein 1 distinguish Epstein-Barr virus strains. Virology 261:79-95. - PubMed
-
- Felsenstein, J. 1981. Evolutionary trees from DNA sequences: a maximum likelihood approach. J. Mol. Evol. 17:368-376. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

