Binding of the 7SK snRNA turns the HEXIM1 protein into a P-TEFb (CDK9/cyclin T) inhibitor
- PMID: 15201869
- PMCID: PMC449783
- DOI: 10.1038/sj.emboj.7600275
Binding of the 7SK snRNA turns the HEXIM1 protein into a P-TEFb (CDK9/cyclin T) inhibitor
Abstract
The positive transcription elongation factor b (P-TEFb) plays a pivotal role in productive elongation of nascent RNA molecules by RNA polymerase II. Core active P-TEFb is composed of CDK9 and cyclin T. In addition, mammalian cell extracts contain an inactive P-TEFb complex composed of four components, CDK9, cyclin T, the 7SK snRNA and the MAQ1/HEXIM1 protein. We now report an in vitro reconstitution of 7SK-dependent HEXIM1 association to purified P-TEFb and subsequent CDK9 inhibition. Yeast three-hybrid tests and gel-shift assays indicated that HEXIM1 binds 7SK snRNA directly and a 7SK snRNA-recognition motif was identified in the central part of HEXIM1 (amino acids (aa) 152-155). Data from yeast two-hybrid and pull-down assay on GST fusion proteins converge to a direct binding of P-TEFb to the HEXIM1 C-terminal domain (aa 181-359). Consistently, point mutations in an evolutionarily conserved motif (aa 202-205) were found to suppress P-TEFb binding and inhibition without affecting 7SK recognition. We propose that the RNA-binding domain of HEXIM1 mediates its association with 7SK and that P-TEFb then enters the complex through association with HEXIM1.
Figures



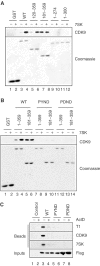





References
-
- Chen R, Yang Z, Zhou Q (2004) Phosphorylated P-TEFb is tagged for inhibition through association with 7SK snRNA. J Biol Chem 279: 4153–4160 - PubMed
-
- Clemens MJ (1997) PKR—a protein kinase regulated by double-stranded RNA. Int J Biochem Cell Biol 29: 945–949 - PubMed
-
- Deleault NR, Lucassen RW, Supattapone S (2003) RNA molecules stimulate prion protein conversion. Nature 425: 717–720 - PubMed
-
- Fraldi A, Licciardo P, Majello B, Giordano A, Lania L (2001) Distinct regions of cyclinT1 are required for binding to CDK9 and for recruitment to the HIV-1 Tat/TAR complex. J Cell Biochem 36: 247–253 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

