Pseudouridine-guide RNAs and other Cbf5p-associated RNAs in Euglena gracilis
- PMID: 15208440
- PMCID: PMC1370595
- DOI: 10.1261/rna.7300804
Pseudouridine-guide RNAs and other Cbf5p-associated RNAs in Euglena gracilis
Abstract
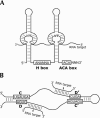
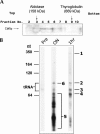
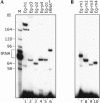
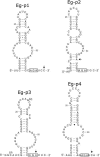
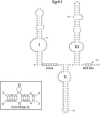
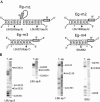
In eukaryotes, box H/ACA small nucleolar RNAs (snoRNAs) guide sites of pseudouridine (Psi) formation in rRNA. These snoRNAs reside in RNP complexes containing the putative Psi synthase, Cbf5p. In this study we have identified Cbf5p-associated RNAs in Euglena gracilis, an early diverging eukaryote, by immunoprecipitating Cbf5p-containing complexes from cellular extracts. We characterized one box H/ACA-like RNA which, however, does not appear to guide Psi formation in rRNA. We also identified four single Psi-guide box AGA RNAs. We determined target sites for these putative Psi-guide RNAs and confirmed that the predicted Psi modifications do, in fact, occur at these positions in Euglena rRNA. The Cbf5p-associated snoRNAs appear to be encoded by multicopy genes, some of which are clustered in the genome together with methylation-guide snoRNA genes. These modification-guide snoRNAs and snoRNA genes are the first ones to be reported in euglenid protists, the evolutionary sister group to the kinetoplastid protozoa. Unexpectedly, we also found and have partially characterized a selenocysteine tRNA homolog in the anti-Cbf5p-immunoprecipitated sample.
Figures



Similar articles
-
Evolutionary appearance of genes encoding proteins associated with box H/ACA snoRNAs: cbf5p in Euglena gracilis, an early diverging eukaryote, and candidate Gar1p and Nop10p homologs in archaebacteria.Nucleic Acids Res. 2000 Jun 15;28(12):2342-52. doi: 10.1093/nar/28.12.2342. Nucleic Acids Res. 2000. PMID: 10871366 Free PMC article.
-
The complete set of H/ACA snoRNAs that guide rRNA pseudouridylations in Saccharomyces cerevisiae.RNA. 2005 Jun;11(6):928-38. doi: 10.1261/rna.2100905. RNA. 2005. PMID: 15923376 Free PMC article.
-
A large collection of compact box C/D snoRNAs and their isoforms in Euglena gracilis: structural, functional and evolutionary insights.J Mol Biol. 2006 Apr 14;357(5):1548-65. doi: 10.1016/j.jmb.2006.01.063. Epub 2006 Feb 8. J Mol Biol. 2006. PMID: 16497322
-
Identifying effects of snoRNA-guided modifications on the synthesis and function of the yeast ribosome.Methods Enzymol. 2007;425:283-316. doi: 10.1016/S0076-6879(07)25013-X. Methods Enzymol. 2007. PMID: 17673089 Review.
-
SnoRNAs as tools for RNA cleavage and modification.Nucleic Acids Symp Ser. 1997;(36):61-3. Nucleic Acids Symp Ser. 1997. PMID: 9478207 Review.
Cited by
-
RNA-Seq employing a novel rRNA depletion strategy reveals a rich repertoire of snoRNAs in Euglena gracilis including box C/D and Ψ-guide RNAs targeting the modification of rRNA extremities.RNA Biol. 2018;15(10):1309-1318. doi: 10.1080/15476286.2018.1526561. Epub 2018 Oct 9. RNA Biol. 2018. PMID: 30252600 Free PMC article.
-
Strong dependence between functional domains in a dual-function snoRNA infers coupling of rRNA processing and modification events.Nucleic Acids Res. 2010 Jun;38(10):3376-87. doi: 10.1093/nar/gkq043. Epub 2010 Feb 9. Nucleic Acids Res. 2010. PMID: 20144950 Free PMC article.
-
Identification of determinants in the protein partners aCBF5 and aNOP10 necessary for the tRNA:Psi55-synthase and RNA-guided RNA:Psi-synthase activities.Nucleic Acids Res. 2007;35(16):5610-24. doi: 10.1093/nar/gkm606. Epub 2007 Aug 17. Nucleic Acids Res. 2007. PMID: 17704128 Free PMC article.
-
Genome-wide analyses of two families of snoRNA genes from Drosophila melanogaster, demonstrating the extensive utilization of introns for coding of snoRNAs.RNA. 2005 Aug;11(8):1303-16. doi: 10.1261/rna.2380905. Epub 2005 Jun 29. RNA. 2005. PMID: 15987805 Free PMC article.
-
Clustered organization, polycistronic transcription, and evolution of modification-guide snoRNA genes in Euglena gracilis.Mol Genet Genomics. 2012 Jan;287(1):55-66. doi: 10.1007/s00438-011-0662-8. Epub 2011 Dec 2. Mol Genet Genomics. 2012. PMID: 22134850
References
-
- Bachellerie, J.-P., Cavaillé, J., and Hüttenhofer, A. 2002. The expanding snoRNA world. Biochimie 84: 775–790. - PubMed
-
- Bakin, A., Lane, B.G., and Ofengand, J. 1994. Clustering of pseudouridine residues around the peptidyltransferase center of yeast cytoplasmic and mitochondrial ribosomes. Biochemistry 33: 13475–13483. - PubMed
-
- Balakin, AG., Smith, L., and Fournier, M.J. 1996. The RNA world of the nucleolus: Two major families of small RNAs defined by different box elements with related functions. Cell 86: 823–834. - PubMed
-
- Barneche, F., Gaspin, C., Guyot R., and Echeverria, M. 2001. Identification of 66 box C/D snoRNAs in Arabidopsis thaliana: Extensive gene duplications generated multiple isoforms predicting new ribosomal RNA 2′-O-methylation sites. J. Mol. Biol. 311: 57–73. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous
