Cystic fibrosis transmembrane conductance regulator degradation depends on the lectins Htm1p/EDEM and the Cdc48 protein complex in yeast
- PMID: 15215312
- PMCID: PMC515346
- DOI: 10.1091/mbc.e04-01-0024
Cystic fibrosis transmembrane conductance regulator degradation depends on the lectins Htm1p/EDEM and the Cdc48 protein complex in yeast
Abstract
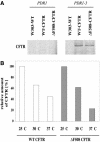
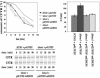
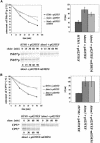
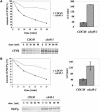
Cystic fibrosis is the most widespread hereditary disease among the white population caused by different mutations of the apical membrane ATP-binding cassette transporter cystic fibrosis transmembrane conductance regulator (CFTR). Its most common mutation, DeltaF508, leads to nearly complete degradation via endoplasmic reticulum-associated degradation (ERAD). Elucidation of the quality control and degradation mechanisms might give rise to new therapeutic approaches to cure this disease. In the yeast Saccharomyces cerevisiae, a variety of components of the protein quality control and degradation system have been identified. Nearly all of these components share homology with mammalian counterparts. We therefore used yeast mutants defective in the ERAD system to identify new components that are involved in human CFTR quality control and degradation. We show the role of the lectin Htm1p in the degradation process of CFTR. Complementation of the HTM1 deficiency in yeast cells by the mammalian orthologue EDEM underlines the necessity of this lectin for CFTR degradation and highlights the similarity of quality control and ERAD in yeast and mammals. Furthermore, degradation of CFTR requires the ubiquitin protein ligases Der3p/Hrd1p and Doa10p as well as the cytosolic trimeric Cdc48p-Ufd1p-Npl4p complex. These proteins also were found to be necessary for ERAD of a mutated yeast "relative" of CFTR, Pdr5(*)p.
Figures


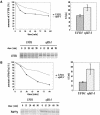
Similar articles
-
Ssz1 restores endoplasmic reticulum-associated protein degradation in cells expressing defective cdc48-ufd1-npl4 complex by upregulating cdc48.Genetics. 2010 Mar;184(3):695-706. doi: 10.1534/genetics.109.111419. Epub 2009 Dec 28. Genetics. 2010. PMID: 20038635 Free PMC article.
-
A Cdc48p-associated factor modulates endoplasmic reticulum-associated degradation, cell stress, and ubiquitinated protein homeostasis.J Biol Chem. 2011 Feb 18;286(7):5744-55. doi: 10.1074/jbc.M110.179259. Epub 2010 Dec 9. J Biol Chem. 2011. PMID: 21148305 Free PMC article.
-
Previously unknown role for the ubiquitin ligase Ubr1 in endoplasmic reticulum-associated protein degradation.Proc Natl Acad Sci U S A. 2013 Sep 17;110(38):15271-6. doi: 10.1073/pnas.1304928110. Epub 2013 Aug 29. Proc Natl Acad Sci U S A. 2013. PMID: 23988329 Free PMC article.
-
Regulation of CFTR Biogenesis by the Proteostatic Network and Pharmacological Modulators.Int J Mol Sci. 2020 Jan 10;21(2):452. doi: 10.3390/ijms21020452. Int J Mol Sci. 2020. PMID: 31936842 Free PMC article. Review.
-
Control of cystic fibrosis transmembrane conductance regulator membrane trafficking: not just from the endoplasmic reticulum to the Golgi.FEBS J. 2013 Sep;280(18):4396-406. doi: 10.1111/febs.12392. Epub 2013 Jul 5. FEBS J. 2013. PMID: 23773658 Review.
Cited by
-
Distinct roles for the Hsp40 and Hsp90 molecular chaperones during cystic fibrosis transmembrane conductance regulator degradation in yeast.Mol Biol Cell. 2004 Nov;15(11):4787-97. doi: 10.1091/mbc.e04-07-0584. Epub 2004 Sep 1. Mol Biol Cell. 2004. PMID: 15342786 Free PMC article.
-
EDEM1's mannosidase-like domain binds ERAD client proteins in a redox-sensitive manner and possesses catalytic activity.J Biol Chem. 2018 Sep 7;293(36):13932-13945. doi: 10.1074/jbc.RA118.004183. Epub 2018 Jul 18. J Biol Chem. 2018. PMID: 30021839 Free PMC article.
-
Targeting ubiquitination machinery in cystic fibrosis: Where do we stand?Cell Mol Life Sci. 2024 Jun 18;81(1):271. doi: 10.1007/s00018-024-05295-z. Cell Mol Life Sci. 2024. PMID: 38888668 Free PMC article. Review.
-
Two salt bridges differentially contribute to the maintenance of cystic fibrosis transmembrane conductance regulator (CFTR) channel function.J Biol Chem. 2013 Jul 12;288(28):20758-67. doi: 10.1074/jbc.M113.476226. Epub 2013 May 24. J Biol Chem. 2013. PMID: 23709221 Free PMC article.
-
The role of the UPS in cystic fibrosis.BMC Biochem. 2007 Nov 22;8 Suppl 1(Suppl 1):S11. doi: 10.1186/1471-2091-8-S1-S11. BMC Biochem. 2007. PMID: 18047735 Free PMC article. Review.
References
-
- Ausubel, F.M., Kingston, R.E., Seidman, F.G., Struhl, K., Moore, D.D., Brent, R., and Smith, F.A. (1992). Current Protocols in Molecular Biology, New York: Greene.
-
- Balzi, E., Wang, M., Leterme, S., Van Dyck, L., and Goffeau, A. (1994). PDR5, a novel yeast multidrug resistance conferring transporter controlled by the transcription regulator PDR1*. J. Biol. Chem. 269, 2206-2214. - PubMed
-
- Bebök, Z., Mazzochi, C., King, S.A., Hong, J.S., and Sorscher, E.J. (1998). The mechanism underlying cystic fibrosis transmembrane conductance regulator transport from the endoplasmic reticulum to the proteasome includes Sec61beta and a cytosolic, deglycosylated intermediary. J. Biol. Chem. 273, 29873-29878. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

