Characterization of a unique group-specific protein (U122) of the severe acute respiratory syndrome coronavirus
- PMID: 15220404
- PMCID: PMC434096
- DOI: 10.1128/JVI.78.14.7311-7318.2004
Characterization of a unique group-specific protein (U122) of the severe acute respiratory syndrome coronavirus
Abstract
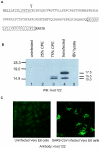
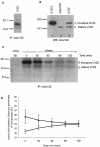
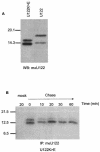
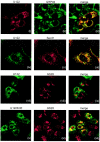
A novel coronavirus (CoV) has been identified as the etiological agent of severe acute respiratory syndrome (SARS). The SARS-CoV genome encodes the characteristic essential CoV replication and structural proteins. Additionally, the genome contains six group-specific open reading frames (ORFs) larger than 50 amino acids, with no known homologues. As with the group-specific genes of the other CoVs, little is known about the SARS-CoV group-specific genes. SARS-CoV ORF7a encodes a putative unique 122-amino-acid protein, designated U122 in this study. The deduced sequence contains a probable cleaved signal sequence and a C-terminal transmembrane helix, indicating that U122 is likely to be a type I membrane protein. The C-terminal tail also contains a typical endoplasmic reticulum (ER) retrieval motif, KRKTE. U122 was expressed in SARS-CoV-infected Vero E6 cells, as it could be detected by Western blot and immunofluorescence analyses. U122 is localized to the perinuclear region of both SARS-CoV-infected and transfected cells and colocalized with ER and intermediate compartment markers. Mutational analyses showed that both the signal peptide sequence and ER retrieval motif were functional.
Figures
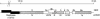
Similar articles
-
A novel severe acute respiratory syndrome coronavirus protein, U274, is transported to the cell surface and undergoes endocytosis.J Virol. 2004 Jul;78(13):6723-34. doi: 10.1128/JVI.78.13.6723-6734.2004. J Virol. 2004. PMID: 15194747 Free PMC article.
-
Structure and intracellular targeting of the SARS-coronavirus Orf7a accessory protein.Structure. 2005 Jan;13(1):75-85. doi: 10.1016/j.str.2004.10.010. Structure. 2005. PMID: 15642263 Free PMC article.
-
Structure, expression, and intracellular localization of the SARS-CoV accessory proteins 7a and 7b.Adv Exp Med Biol. 2006;581:115-20. doi: 10.1007/978-0-387-33012-9_20. Adv Exp Med Biol. 2006. PMID: 17037516 Free PMC article. No abstract available.
-
Understanding the accessory viral proteins unique to the severe acute respiratory syndrome (SARS) coronavirus.Antiviral Res. 2006 Nov;72(2):78-88. doi: 10.1016/j.antiviral.2006.05.010. Epub 2006 Jun 6. Antiviral Res. 2006. PMID: 16820226 Free PMC article. Review.
-
Characterization of viral proteins encoded by the SARS-coronavirus genome.Antiviral Res. 2005 Feb;65(2):69-78. doi: 10.1016/j.antiviral.2004.10.001. Antiviral Res. 2005. PMID: 15708633 Free PMC article. Review.
Cited by
-
Pathology of the thyroid in severe acute respiratory syndrome.Hum Pathol. 2007 Jan;38(1):95-102. doi: 10.1016/j.humpath.2006.06.011. Epub 2006 Sep 25. Hum Pathol. 2007. PMID: 16996569 Free PMC article.
-
Induction of apoptosis by the severe acute respiratory syndrome coronavirus 7a protein is dependent on its interaction with the Bcl-XL protein.J Virol. 2007 Jun;81(12):6346-55. doi: 10.1128/JVI.00090-07. Epub 2007 Apr 11. J Virol. 2007. PMID: 17428862 Free PMC article.
-
A SARS-CoV protein, ORF-6, induces caspase-3 mediated, ER stress and JNK-dependent apoptosis.Biochim Biophys Acta. 2008 Dec;1780(12):1383-7. doi: 10.1016/j.bbagen.2008.07.009. Epub 2008 Jul 28. Biochim Biophys Acta. 2008. PMID: 18708124 Free PMC article.
-
SARS-CoV-2 accessory proteins ORF7a and ORF3a use distinct mechanisms to down-regulate MHC-I surface expression.Proc Natl Acad Sci U S A. 2023 Jan 3;120(1):e2208525120. doi: 10.1073/pnas.2208525120. Epub 2022 Dec 27. Proc Natl Acad Sci U S A. 2023. PMID: 36574644 Free PMC article.
-
Object-oriented biological system integration: a SARS coronavirus example.Bioinformatics. 2005 May 15;21(10):2502-9. doi: 10.1093/bioinformatics/bti344. Epub 2005 Feb 24. Bioinformatics. 2005. PMID: 15731211 Free PMC article.
References
-
- Anand, K., J. Ziebuhr, P. Wadhwani, J. R. Mesters, and R. Hilgenfeld. 2003. Coronavirus main proteinase (3CLpro) structure: basis for design of anti-SARS drugs. Science 300:1763-1767. - PubMed
-
- Eichler, R., O. Lenz, T. Strecker, and W. Garten. 2003. Signal peptide of Lassa virus glycoproteins GP-C exhibits an unusual length. FEBS Lett. 538:203-206. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

