Two breakpoint clusters at fragile site FRA3B form phased nucleosomes
- PMID: 15231750
- PMCID: PMC442151
- DOI: 10.1101/gr.2304404
Two breakpoint clusters at fragile site FRA3B form phased nucleosomes
Abstract
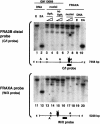
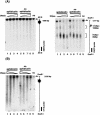
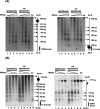
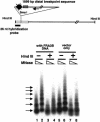
Fragile sites are gaps and breaks in metaphase chromosomes generated by specific culture conditions. Fragile site FRA3B is the most unstable site and is directly involved in the breakpoints of deletion and translocation in a wide spectrum of cancers. To learn about the general characteristics of common fragile sites, we investigated the chromatin structure of the FRA3B site. Because FRA3B spans several hundred kilobases, we focused our study on two breakpoint clusters found in FRA3B. Using various nucleases, we demonstrated that these two regions contain phased nucleosomes, regardless of treatment with aphidicolin. Because these regions are located in intron 4 of the FHIT gene, it is very interesting to observe phased nucleosomes over these regions, which are several hundred kilobases downstream from the promoter. Further, by using nucleosome assembly assays, we demonstrate that these two regions do not contain strong nucleosome positioning elements. These results suggest that other factors appear to cooperate with the DNA sequence of these regions to impart nucleosome phasing. This study provides the first information on the chromatin structure of breakpoint regions in a common fragile site. The observation of phased nucleosomes over these breakpoint regions could offer a foundation to understand the expression of fragile sites.
Copyright 2004 Cold Spring Harbor Laboratory Press ISSN
Figures

Similar articles
-
Identification of unstable sequences within the common fragile site at 3p14.2: implications for the mechanism of deletions within fragile histidine triad gene/common fragile site at 3p14.2 in tumors.Cancer Res. 2002 Jun 15;62(12):3477-84. Cancer Res. 2002. PMID: 12067991
-
Positions of chromosome 3p14.2 fragile sites (FRA3B) within the FHIT gene.Cancer Res. 1997 Mar 15;57(6):1166-70. Cancer Res. 1997. PMID: 9067288
-
Translocation breakpoints in FHIT and FRA3B in both homologs of chromosome 3 in an esophageal adenocarcinoma.Genes Chromosomes Cancer. 2001 Mar;30(3):292-8. doi: 10.1002/1098-2264(2000)9999:9999<::aid-gcc1095>3.0.co;2-f. Genes Chromosomes Cancer. 2001. PMID: 11170287
-
Common fragile sites and cancer (review).Int J Oncol. 1998 Jan;12(1):187-96. Int J Oncol. 1998. PMID: 9454904 Review.
-
The role of the FHIT/FRA3B locus in cancer.Annu Rev Genet. 1998;32:7-31. doi: 10.1146/annurev.genet.32.1.7. Annu Rev Genet. 1998. PMID: 9928473 Review.
Cited by
-
Allelic imbalance and abnormal expression of FHIT in endemic nasopharyngeal carcinoma: association with clinicopathological features.Eur Arch Otorhinolaryngol. 2010 Dec;267(12):1933-41. doi: 10.1007/s00405-010-1301-4. Epub 2010 Jun 15. Eur Arch Otorhinolaryngol. 2010. PMID: 20552362
-
DNA topoisomerases participate in fragility of the oncogene RET.PLoS One. 2013 Sep 11;8(9):e75741. doi: 10.1371/journal.pone.0075741. eCollection 2013. PLoS One. 2013. PMID: 24040417 Free PMC article.
-
Fragile histidine triad protein: structure, function, and its association with tumorogenesis.J Cancer Res Clin Oncol. 2010 Mar;136(3):333-50. doi: 10.1007/s00432-009-0751-9. Epub 2009 Dec 24. J Cancer Res Clin Oncol. 2010. PMID: 20033706 Free PMC article. Review.
-
Are common fragile sites merely structural domains or highly organized "functional" units susceptible to oncogenic stress?Cell Mol Life Sci. 2014 Dec;71(23):4519-44. doi: 10.1007/s00018-014-1717-x. Epub 2014 Sep 20. Cell Mol Life Sci. 2014. PMID: 25238782 Free PMC article. Review.
-
Analysis of the t(3;8) of hereditary renal cell carcinoma: a palindrome-mediated translocation.Cancer Genet. 2014 Apr;207(4):133-40. doi: 10.1016/j.cancergen.2014.03.004. Epub 2014 Mar 18. Cancer Genet. 2014. PMID: 24813807 Free PMC article.
References
-
- Boldog, F., Gemmill, R.M., West, J., Robinson, M., Robinson, L., Li, E., Roche, J., Todd, S., Waggoner, B., Lundstrom, R., et al. 1997. Chromosome 3p14 homozygous deletions and sequence analysis of FRA3B. Hum. Mol. Genet. 6: 193–203. - PubMed
-
- Casper, A.M., Nghiem, P., Arlt, M.F., and Glover, T.W. 2002. ATR regulates fragile site stability. Cell 111: 779–789. - PubMed
-
- Corbin, S., Neilly, M.E., Espinosa III, R., Davis, E.M., McKeithan, T.W., and Le Beau, M.M. 2002. Identification of unstable sequences within the common fragile site at 3p14.2: Implications for the mechanism of deletions within fragile histidine triad gene/common fragile site at 3p14.2 in tumors. Cancer Res. 62: 3477–3484. - PubMed
-
- Eberhart, D.E. and Warren, S.T. 1996. Nuclease sensitivity of permeabilized cells confirms altered chromatin formation at the fragile X locus. Somat. Cell Mol. Genet. 22: 435–441. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
