Use of oligonucleotide microarrays for rapid detection and serotyping of acute respiratory disease-associated adenoviruses
- PMID: 15243087
- PMCID: PMC446281
- DOI: 10.1128/JCM.42.7.3232-3239.2004
Use of oligonucleotide microarrays for rapid detection and serotyping of acute respiratory disease-associated adenoviruses
Abstract
The cessation of the adenovirus vaccination program for military trainees has resulted in several recent acute respiratory disease (ARD) outbreaks. In the absence of vaccination, rapid detection methods are necessary for the timely implementation of measures to prevent adenovirus transmission within military training facilities. To this end, we have combined a fluorogenic real-time multiplex PCR assay with four sets of degenerate PCR primers that target the E1A, fiber, and hexon genes with a long oligonucleotide microarray capable of identifying the most common adenovirus serotypes associated with adult respiratory tract infections (serotypes 3, 4, 7, 16, and 21) and a representative member of adenovirus subgroup C (serotype 6) that is a common cause of childhood ARD and that often persists into adulthood. Analyses with prototype strains demonstrated unique hybridization patterns for representative members of adenovirus subgroups B(1), B(2), C, and E, thus allowing serotype determination. Microarray-based sensitivity assessments revealed lower detection limits (between 1 and 100 genomic copies) for adenovirus serotype 4 (Ad4) and Ad7 cell culture lysates, clinical nasal washes, and throat swabs and purified DNA from clinical samples. When adenovirus was detected from coded clinical samples, the results obtained by this approach demonstrated an excellent concordance with those obtained by the more established method of adenovirus identification as well as by cell culture with fluorescent-antibody staining. Finally, the utility of this method was further supported by its ability to detect adenoviral coinfections, contamination, and, potentially, recombination events. Taken together, the results demonstrate the usefulness of the simple and rapid diagnostic method developed for the unequivocal identification of ARD-associated adenoviral serotypes from laboratory or clinical samples that can be completed in 1.5 to 4.0 h.
Figures



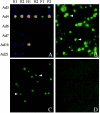
References
-
- Avellón, A., P. Perez, J. C. Aguilar, R. Lejarazu, and J. E. Echevarria. 2001. Rapid and sensitive diagnosis of human adenovirus infections by a generic polymerase chain reaction. J. Virol. Methods 92:113-120. - PubMed
-
- Charles, P. T., G. J. Vora, J. D. Andreadis, A. J. Fortney, C. E. Meador, C. S. Dulcey, and D. A. Stenger. 2003. Fabrication and surface characterization of DNA microarrays using amine- and thiol-terminated oligonucleotide probes. Langmuir 19:1586-1591.
-
- De Jong, J. C., A. G. Wermenbol, M. W. Verweij-Uijterwaal, K. W. Slaterus, P. Wertheim-Van Dillen, G. J. J. Van Doornum, S. H. Khoo, and J. C. Hierholzer. 1999. Adenoviruses from human immunodeficiency virus-infected individuals, including two strains that represent new candidate serotypes Ad50 and Ad51 of species B1 and D, respectively. J. Clin. Microbiol. 37:3940-3945. - PMC - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

