High genetic diversity revealed by variable-number tandem repeat genotyping and analysis of hsp65 gene polymorphism in a large collection of "Mycobacterium canettii" strains indicates that the M. tuberculosis complex is a recently emerged clone of "M. canettii"
- PMID: 15243089
- PMCID: PMC446256
- DOI: 10.1128/JCM.42.7.3248-3255.2004
High genetic diversity revealed by variable-number tandem repeat genotyping and analysis of hsp65 gene polymorphism in a large collection of "Mycobacterium canettii" strains indicates that the M. tuberculosis complex is a recently emerged clone of "M. canettii"
Abstract
We have analyzed, using complementary molecular methods, the diversity of 43 strains of "Mycobacterium canettii" originating from the Republic of Djibouti, on the Horn of Africa, from 1998 to 2003. Genotyping by multiple-locus variable-number tandem repeat analysis shows that all the strains belong to a single but very distant group when compared to strains of the Mycobacterium tuberculosis complex (MTBC). Thirty-one strains cluster into one large group with little variability and five strains form another group, whereas the other seven are more diverged. In total, 14 genotypes are observed. The DR locus analysis reveals additional variability, some strains being devoid of a direct repeat locus and others having unique spacers. The hsp65 gene polymorphism was investigated by restriction enzyme analysis and sequencing of PCR amplicons. Four new single nucleotide polymorphisms were discovered. One strain was characterized by three nucleotide changes in 441 bp, creating new restriction enzyme polymorphisms. As no sequence variability was found for hsp65 in the whole MTBC, and as a single point mutation separates M. tuberculosis from the closest "M. canettii" strains, this diversity within "M. canettii" subspecies strongly suggests that it is the most probable source species of the MTBC rather than just another branch of the MTBC.
Figures



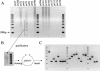

References
-
- Brosch, R., S. V. Gordon, M. Marmiesse, P. Brodin, C. Buchrieser, K. Eiglmeier, T. Garnier, C. Gutierrez, G. Hewinson, K. Kremer, L. M. Parsons, A. S. Pym, S. Samper, D. van Soolingen, and S. T. Cole. 2002. A new evolutionary scenario for the Mycobacterium tuberculosis complex. Proc. Natl. Acad. Sci. USA 99:3684-3689. - PMC - PubMed
-
- Embley, T. M. 1991. The linear PCR reaction: a simple and robust method for sequencing amplified rRNA genes. Lett. Appl. Microbiol. 13:171-174. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

