Extending the classification of bacterial transcription factors beyond the helix-turn-helix motif as an alternative approach to discover new cis/trans relationships
- PMID: 15247334
- PMCID: PMC443547
- DOI: 10.1093/nar/gkh673
Extending the classification of bacterial transcription factors beyond the helix-turn-helix motif as an alternative approach to discover new cis/trans relationships
Abstract
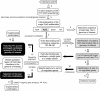
Transcription factors (TFs) of bacterial helix-turn-helix superfamilies exhibit different effector-binding domains (EBDs) fused to a DNA-binding domain with a common feature. In a previous study of the GntR superfamily, we demonstrated that classifying members into subfamilies according to the EBD heterogeneity highlighted unsuspected and accurate TF-binding site signatures. In this work, we present how such in silico analysis can provide prediction tools to discover new cis/trans relationships. The TF-binding site consensus of the HutC/GntR subfamily was used to (i) predict target sites within the Streptomyces coelicolor genome, (ii) discover a new HutC/GntR regulon and (iii) discover its specific TF. By scanning the S.coelicolor genome we identified a presumed new HutC regulon that comprises genes of the phosphotransferase system (PTS) specific for the uptake of N-acetylglucosamine (PTS(Nag)). A weight matrix was derived from the compilation of the predicted cis-acting elements upstream of each gene of the presumed regulon. Under the assumption that TFs are often subject to autoregulation, we used this matrix to scan the upstream region of the 24 HutC-like members of S.coelicolor. orf SCO5231 (dasR) was selected as the best candidate according to the high score of a 16 bp sequence identified in its upstream region. Our prediction that DasR regulates the PTS(Nag) regulon was confirmed by in vivo and in vitro experiments. In conclusion, our in silico approach permitted to highlight the specific TF of a regulon out of the 673 orfs annotated as 'regulatory proteins' within the genome of S.coelicolor.
Figures
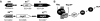

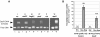
Similar articles
-
The sugar phosphotransferase system of Streptomyces coelicolor is regulated by the GntR-family regulator DasR and links N-acetylglucosamine metabolism to the control of development.Mol Microbiol. 2006 Sep;61(5):1237-51. doi: 10.1111/j.1365-2958.2006.05319.x. Mol Microbiol. 2006. PMID: 16925557
-
Crystal Structures of the Global Regulator DasR from Streptomyces coelicolor: Implications for the Allosteric Regulation of GntR/HutC Repressors.PLoS One. 2016 Jun 23;11(6):e0157691. doi: 10.1371/journal.pone.0157691. eCollection 2016. PLoS One. 2016. PMID: 27337024 Free PMC article.
-
GntR Family of Bacterial Transcription Factors and Their DNA Binding Motifs: Structure, Positioning and Co-Evolution.PLoS One. 2015 Jul 7;10(7):e0132618. doi: 10.1371/journal.pone.0132618. eCollection 2015. PLoS One. 2015. PMID: 26151451 Free PMC article.
-
The many faces of the helix-turn-helix domain: transcription regulation and beyond.FEMS Microbiol Rev. 2005 Apr;29(2):231-62. doi: 10.1016/j.femsre.2004.12.008. FEMS Microbiol Rev. 2005. PMID: 15808743 Review.
-
Chapter 1: Variation in form and function the helix-turn-helix regulators of the GntR superfamily.Adv Appl Microbiol. 2009;69:1-22. doi: 10.1016/S0065-2164(09)69001-8. Adv Appl Microbiol. 2009. PMID: 19729089 Review.
Cited by
-
Importance of the 5' regulatory region to bacterial synthetic biology applications.Microb Biotechnol. 2021 Nov;14(6):2291-2315. doi: 10.1111/1751-7915.13868. Epub 2021 Jun 25. Microb Biotechnol. 2021. PMID: 34171170 Free PMC article. Review.
-
A single transcription factor regulates evolutionarily diverse but functionally linked metabolic pathways in response to nutrient availability.Mol Syst Biol. 2009;5:282. doi: 10.1038/msb.2009.40. Epub 2009 Jun 16. Mol Syst Biol. 2009. PMID: 19536205 Free PMC article.
-
Allosteric regulation by c-di-AMP modulates a complete N-acetylglucosamine signaling cascade in Saccharopolyspora erythraea.Nat Commun. 2024 May 7;15(1):3825. doi: 10.1038/s41467-024-48063-0. Nat Commun. 2024. PMID: 38714645 Free PMC article.
-
Functional analysis of the N-acetylglucosamine metabolic genes of Streptomyces coelicolor and role in control of development and antibiotic production.J Bacteriol. 2012 Mar;194(5):1136-44. doi: 10.1128/JB.06370-11. Epub 2011 Dec 22. J Bacteriol. 2012. PMID: 22194457 Free PMC article.
-
Discovery of a Cryptic Antifungal Compound from Streptomyces albus J1074 Using High-Throughput Elicitor Screens.J Am Chem Soc. 2017 Jul 12;139(27):9203-9212. doi: 10.1021/jacs.7b02716. Epub 2017 Jun 29. J Am Chem Soc. 2017. PMID: 28590725 Free PMC article.
References
-
- Pabo C.O. and Sauer,R.T. (1992) Transcription factors: structural families and principles of DNA recognition. Annu. Rev. Biochem., 61, 1053–1095. - PubMed
-
- Rosinski J.A. and Atchley,W.R. (1999) Molecular evolution of helix–turn–helix proteins. J. Mol. Evol., 49, 301–309. - PubMed
-
- Brown N.L., Stoyanov,J.V., Kidd,S.P. and Hobman,J.L. (2003) The MerR family of transcriptional regulators. FEMS Microbiol. Rev., 27,145–163. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

