A phage display selection of engrailed homeodomain mutants and the importance of residue Q50
- PMID: 15247345
- PMCID: PMC484177
- DOI: 10.1093/nar/gkh690
A phage display selection of engrailed homeodomain mutants and the importance of residue Q50
Abstract
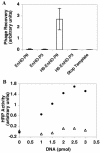
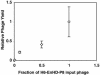
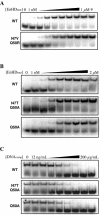
Mutants of engrailed homeodomain (HD) that retain DNA-binding activity were isolated using a phage display selection. This selection was used to enrich for active DNA-binding clones from a complex library consisting of over a billion members. A more focused library of mutant homeodomains consisting of all possible amino acid combinations at two DNA-contacting residues (I47 and Q50) was constructed and screened for members capable of binding tightly and specifically to the engrailed consensus sequence, TAATTA. The isolated mutants largely recapitulated the distribution of amino acids found at these positions in natural homeodomains thus validating the in vitro selection conditions. In particular, the unequivocal advantage enjoyed by glutamine at residue 50 is surprising in light of reports that minimize the importance of this residue. Here, the subtle contributions of residue Q50 are demonstrated to play a functionally important role in specific recognition of DNA. These results highlight the complex subtlety of protein-DNA interactions, underscoring the value of the first reported in vitro selection of a homeodomain.
Figures
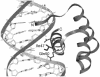

Similar articles
-
Structure and properties of a re-engineered homeodomain protein-DNA interface.ACS Chem Biol. 2006 Dec 15;1(12):755-60. doi: 10.1021/cb6003756. ACS Chem Biol. 2006. PMID: 17240973
-
Adaptability at a protein-DNA interface: re-engineering the engrailed homeodomain to recognize an unnatural nucleotide.J Am Chem Soc. 2004 Jul 7;126(26):8078-9. doi: 10.1021/ja048113w. J Am Chem Soc. 2004. PMID: 15225024
-
Differential DNA-binding specificity of the engrailed homeodomain: the role of residue 50.Biochemistry. 1994 Aug 9;33(31):9187-94. doi: 10.1021/bi00197a022. Biochemistry. 1994. PMID: 8049221
-
Mutational analysis of the engrailed homeodomain recognition helix by phage display.Nucleic Acids Res. 1999 Feb 15;27(4):1182-9. doi: 10.1093/nar/27.4.1182. Nucleic Acids Res. 1999. PMID: 9927754 Free PMC article.
-
Determinants of the DNA-binding specificity of the Avian homeodomain protein, AKR.DNA Cell Biol. 1999 Oct;18(10):791-804. doi: 10.1089/104454999314935. DNA Cell Biol. 1999. PMID: 10541438
Cited by
-
Diversifying the structure of zinc finger nucleases for high-precision genome editing.Nat Commun. 2019 Mar 8;10(1):1133. doi: 10.1038/s41467-019-08867-x. Nat Commun. 2019. PMID: 30850604 Free PMC article.
-
Profiling the DNA-binding specificities of engineered Cys2His2 zinc finger domains using a rapid cell-based method.Nucleic Acids Res. 2007;35(11):e81. doi: 10.1093/nar/gkm385. Epub 2007 May 30. Nucleic Acids Res. 2007. PMID: 17537811 Free PMC article.
-
Sequence specificity of single-stranded DNA-binding proteins: a novel DNA microarray approach.Nucleic Acids Res. 2007;35(10):e75. doi: 10.1093/nar/gkm040. Epub 2007 May 8. Nucleic Acids Res. 2007. PMID: 17488853 Free PMC article.
-
Reprogramming cells with synthetic proteins.Asian J Androl. 2015 May-Jun;17(3):394-402. doi: 10.4103/1008-682X.145433. Asian J Androl. 2015. PMID: 25652623 Free PMC article. Review.
-
Probing the informational and regulatory plasticity of a transcription factor DNA-binding domain.PLoS Genet. 2012;8(3):e1002614. doi: 10.1371/journal.pgen.1002614. Epub 2012 Mar 29. PLoS Genet. 2012. PMID: 22496663 Free PMC article.
References
-
- Pabo C.O. and Nekludova,L. (2000) Geometric analysis and comparison of protein–DNA interfaces: why is there no simple code for recognition? J. Mol. Biol., 301, 597–624. - PubMed
-
- Pabo C.O., Peisach,E. and Grant,R.A. (2001) Design and selection of novel Cys(2)His(2) zinc finger proteins. Annu. Rev. Biochem., 70, 313–340. - PubMed
-
- Greisman H.A. and Pabo,C.O. (1997) A general strategy for selecting high-affinity zinc finger proteins for diverse DNA target sites. Science, 275, 657–661. - PubMed

