Overexpression of OsRAA1 causes pleiotropic phenotypes in transgenic rice plants, including altered leaf, flower, and root development and root response to gravity
- PMID: 15247372
- PMCID: PMC519066
- DOI: 10.1104/pp.104.041996
Overexpression of OsRAA1 causes pleiotropic phenotypes in transgenic rice plants, including altered leaf, flower, and root development and root response to gravity
Abstract
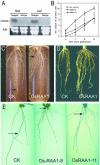
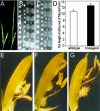
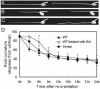
There are very few root genes that have been described in rice as a monocotyledonous model plant so far. Here, the OsRAA1 (Oryza sativa Root Architecture Associated 1) gene has been characterized molecularly. OsRAA1 encodes a 12.0-kD protein that has 58% homology to the AtFPF1 (Flowering Promoting Factor 1) in Arabidopsis, which has not been reported as modulating root development yet. Data of in situ hybridization and OsRAA1::GUS transgenic plant showed that OsRAA1 expressed specifically in the apical meristem, the elongation zone of root tip, steles of the branch zone, and the young lateral root. Constitutive expression of OsRAA1 under the control of maize (Zea mays) ubiquitin promoter resulted in phenotypes of reduced growth of primary root, increased number of adventitious roots and helix primary root, and delayed gravitropic response of roots in seedlings of rice (Oryza sativa), which are similar to the phenotypes of the wild-type plant treated with auxin. With overexpression of OsRAA1, initiation and growth of adventitious root were more sensitive to treatment of auxin than those of the control plants, while their responses to 9-hydroxyfluorene-9-carboxylic acid in both transgenic line and wild type showed similar results. OsRAA1 constitutive expression also caused longer leaves and sterile florets at the last stage of plant development. Analysis of northern blot and GUS activity staining of OsRAA1::GUS transgenic plants demonstrated that the OsRAA1 expression was induced by auxin. At the same time, overexpression of OsRAA1 also caused endogenous indole-3-acetic acid to increase. These data suggested that OsRAA1 as a new gene functions in the development of rice root systems, which are mediated by auxin. A positive feedback regulation mechanism of OsRAA1 to indole-3-acetic acid metabolism may be involved in rice root development in nature.
Figures






Similar articles
-
Rice ROOT ARCHITECTURE ASSOCIATED1 binds the proteasome subunit RPT4 and is degraded in a D-box and proteasome-dependent manner.Plant Physiol. 2008 Oct;148(2):843-55. doi: 10.1104/pp.108.125294. Epub 2008 Aug 13. Plant Physiol. 2008. PMID: 18701670 Free PMC article.
-
OsAUX1 controls lateral root initiation in rice (Oryza sativa L.).Plant Cell Environ. 2015 Nov;38(11):2208-22. doi: 10.1111/pce.12467. Epub 2015 Jan 7. Plant Cell Environ. 2015. PMID: 25311360
-
FPF1 transgene leads to altered flowering time and root development in rice.Plant Cell Rep. 2005 May;24(2):79-85. doi: 10.1007/s00299-004-0906-8. Epub 2005 Jan 22. Plant Cell Rep. 2005. PMID: 15666165
-
APC-targeted RAA1 degradation mediates the cell cycle and root development in plants.Plant Signal Behav. 2010 Mar;5(3):218-23. doi: 10.4161/psb.5.3.10661. Epub 2010 Mar 14. Plant Signal Behav. 2010. PMID: 20037474 Free PMC article. Review.
-
Genetic and Hormonal Blueprint of Shoot-Borne Adventitious Root Development in Rice and Maize.Plant Cell Physiol. 2023 Jan 30;63(12):1806-1813. doi: 10.1093/pcp/pcac084. Plant Cell Physiol. 2023. PMID: 35713294 Review.
Cited by
-
ODDSOC2 is a MADS box floral repressor that is down-regulated by vernalization in temperate cereals.Plant Physiol. 2010 Jul;153(3):1062-73. doi: 10.1104/pp.109.152488. Epub 2010 Apr 29. Plant Physiol. 2010. PMID: 20431086 Free PMC article.
-
OsAPL controls the nutrient transport systems in the leaf of rice (Oryza sativa L.).Planta. 2022 Jun 14;256(1):11. doi: 10.1007/s00425-022-03913-3. Planta. 2022. PMID: 35699777
-
Upland cotton gene GhFPF1 confers promotion of flowering time and shade-avoidance responses in Arabidopsis thaliana.PLoS One. 2014 Mar 13;9(3):e91869. doi: 10.1371/journal.pone.0091869. eCollection 2014. PLoS One. 2014. PMID: 24626476 Free PMC article.
-
Importance of pre-mRNA splicing and its study tools in plants.Adv Biotechnol (Singap). 2024 Feb 8;2(1):4. doi: 10.1007/s44307-024-00009-9. Adv Biotechnol (Singap). 2024. PMID: 39883322 Free PMC article. Review.
-
Genome-wide meta-analysis of QTL for morphological related traits of flag leaf in bread wheat.PLoS One. 2022 Oct 24;17(10):e0276602. doi: 10.1371/journal.pone.0276602. eCollection 2022. PLoS One. 2022. PMID: 36279291 Free PMC article.
References
-
- Bernasconi P (1998) Auxin. In L Taiz, E Zeiger, eds, Plant Physiology. Sinauer Associates, Sunderland, MA, pp 543–590
-
- Bleecker A, Schuette JL, Kende H (1986) Anatomical analysis of growth and developmental patterns in the internode of deepwater rice. Planta 169: 490–497 - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

