Abscisic acid induces CBF gene transcription and subsequent induction of cold-regulated genes via the CRT promoter element
- PMID: 15247382
- PMCID: PMC519084
- DOI: 10.1104/pp.104.043562
Abscisic acid induces CBF gene transcription and subsequent induction of cold-regulated genes via the CRT promoter element
Abstract
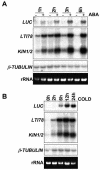
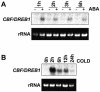
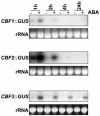
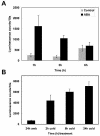
Many cold-regulated genes of Arabidopsis are inducible by abscisic acid (ABA) as well as by cold. This has been thought to occur via two separate signaling pathways, with ABA acting via ABA-responsive promoter elements and low temperature activating the C-repeat element (CRT; dehydration-responsive) promoter element via CBF (DREB1) transcription factors. We show here that ABA is also capable of activating the CRT promoter element. Although the more recently discovered ABA-inducible CBF4 transcription factor might have accounted for this, we show here that CBF1-3 transcript levels also increase in response to elevated ABA levels. This increase in CBF1-3 transcript levels appears to be at least in part due to increased activity of the CBF promoters in response to ABA. A total of 125 bp of the CBF2 promoter, which has previously been shown to be sufficient for cold-, mechanical-, and cycloheximide-induced expression, was also sufficient for ABA-induced expression. However, the ABA-responsive promoter element-like motif within this region is not needed for ABA-induced expression. An observed increase in CBF protein levels after ABA treatment, together with previous data showing that increased CBF levels are sufficient for cold-regulated gene induction, suggests that ABA-induced increases in CBF1-3 transcript levels do have the potential to activate the CRT. Our data indicate therefore that activation of the CRT may also occur via a novel ABA-inducible signaling pathway using the normally cold-inducible CBFs.
Figures


Similar articles
-
Quantitative statistical analysis of cis-regulatory sequences in ABA/VP1- and CBF/DREB1-regulated genes of Arabidopsis.Plant Physiol. 2005 Sep;139(1):437-47. doi: 10.1104/pp.104.058412. Epub 2005 Aug 19. Plant Physiol. 2005. PMID: 16113229 Free PMC article.
-
Combinatorial interactions of multiple cis-elements regulating the induction of the Arabidopsis XERO2 dehydrin gene by abscisic acid and cold.Plant J. 2008 Apr;54(1):15-29. doi: 10.1111/j.1365-313X.2007.03399.x. Epub 2007 Dec 15. Plant J. 2008. PMID: 18088305
-
Low temperature regulation of the Arabidopsis CBF family of AP2 transcriptional activators as an early step in cold-induced COR gene expression.Plant J. 1998 Nov;16(4):433-42. doi: 10.1046/j.1365-313x.1998.00310.x. Plant J. 1998. PMID: 9881163
-
CBF-dependent signaling pathway: a key responder to low temperature stress in plants.Crit Rev Biotechnol. 2011 Jun;31(2):186-92. doi: 10.3109/07388551.2010.505910. Epub 2010 Oct 4. Crit Rev Biotechnol. 2011. PMID: 20919819 Review.
-
Molecular switch for cold acclimation -- anatomy of the cold-inducible promoter in plants.Biochemistry (Mosc). 2013 Apr;78(4):342-54. doi: 10.1134/S0006297913040032. Biochemistry (Mosc). 2013. PMID: 23590437 Review.
Cited by
-
Expression analysis and functional characterization of a novel cold-responsive gene CbCOR15a from Capsella bursa-pastoris.Mol Biol Rep. 2012 May;39(5):5169-79. doi: 10.1007/s11033-011-1313-1. Epub 2011 Dec 9. Mol Biol Rep. 2012. PMID: 22160575
-
Consensus by democracy. Using meta-analyses of microarray and genomic data to model the cold acclimation signaling pathway in Arabidopsis.Plant Physiol. 2006 Aug;141(4):1219-32. doi: 10.1104/pp.106.083527. Plant Physiol. 2006. PMID: 16896234 Free PMC article.
-
Hormonal Orchestration of Bud Dormancy Cycle in Deciduous Woody Perennials.Front Plant Sci. 2019 Sep 18;10:1136. doi: 10.3389/fpls.2019.01136. eCollection 2019. Front Plant Sci. 2019. PMID: 31620159 Free PMC article. Review.
-
Identification of genes associated with the regulation of cold tolerance and the RNA movement in the grafted apple.Sci Rep. 2023 Jul 18;13(1):11583. doi: 10.1038/s41598-023-38571-2. Sci Rep. 2023. PMID: 37463950 Free PMC article.
-
A Glycine-Rich RNA-Binding Protein, CsGR-RBP3, Is Involved in Defense Responses Against Cold Stress in Harvested Cucumber (Cucumis sativus L.) Fruit.Front Plant Sci. 2018 Apr 23;9:540. doi: 10.3389/fpls.2018.00540. eCollection 2018. Front Plant Sci. 2018. PMID: 29740470 Free PMC article.
References
-
- Baker SS, Wilhelm KS, Thomashow MF (1994) The 5′-region of Arabidopsis thaliana cor15a has cis-acting elements that confer cold-, drought- and ABA-regulated gene expression. Plant Mol Biol 24: 701–713 - PubMed
-
- Boyce JM, Knight H, Deyholos M, Openshaw MR, Galbraith DW, Warren G, Knight MR (2003) The sfr6 mutant of Arabidopsis is defective in transcriptional activation via CBF/DREB1 and DREB2 and shows sensitivity to osmotic stress. Plant J 34: 395–406 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

