Measurement of DNA mismatch repair activity in live cells
- PMID: 15249596
- PMCID: PMC484197
- DOI: 10.1093/nar/gnh098
Measurement of DNA mismatch repair activity in live cells
Abstract
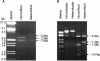
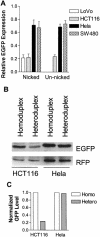
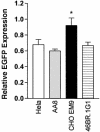
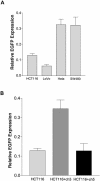
Loss of DNA mismatch repair (MMR) function leads to the development and progression of certain cancers. Currently, assays for DNA MMR activity involve the use of cell extracts and are technically challenging and costly. Here, we report a rapid, less labor-intensive method that can quantitatively measure MMR activity in live cells. A G-G or T-G mismatch was introduced into the ATG start codon of the enhanced green fluorescent protein (EGFP) gene. Repair of the G-G or T-G mismatch to G-C or T-A, respectively, in the heteroduplex plasmid generates a functional EGFP gene expression. The heteroduplex plasmid and a similarly constructed homoduplex plasmid were transfected in parallel into the same cell line and the number of green cells counted by flow cytometry. Relative EGFP expression was calculated as the total fluorescence intensity of cells transfected with the heteroduplex construct divided by that of cells transfected with the homoduplex construct. We have tested several cell lines from both MMR-deficient and MMR-proficient groups using this method, including a colon carcinoma cell line HCT116 with defective hMLH1 gene and a derivative complemented by transient transfection with hMLH1 cDNA. Results show that MMR-proficient cells have significantly higher EGFP expression than MMR-deficient cells, and that transient expression of hMLH1 alone can elevate MMR activity in HCT116 cells. This method is potentially useful in comparing and monitoring MMR activity in live cells under various growth conditions.
Figures


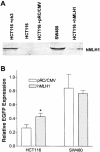
References
-
- Modrich P. and Lahue,R. (1996) Mismatch repair in replication fidelity, genetic recombination, and cancer biology. Annu. Rev. Biochem., 65, 101–133. - PubMed
-
- Fishel R. (1998) Mismatch repair, molecular switches, and signal transduction. Genes Dev., 12, 2096–2101. - PubMed
-
- Umar A. and Kunkel,T.A. (1996) DNA-replication fidelity, mismatch repair and genome instability in cancer cells. Eur. J. Biochem., 238, 297–307. - PubMed
-
- Kolodner R.D. and Marsischky,G.T. (1999) Eukaryotic DNA mismatch repair. Curr. Opin. Genet. Dev., 9, 89–96. - PubMed
-
- Modrich P. (1997) Strand-specific mismatch repair in mammalian cells. J. Biol. Chem., 272, 24727–24730. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

