Intracellular processing of human herpesvirus 6 glycoproteins Q1 and Q2 into tetrameric complexes expressed on the viral envelope
- PMID: 15254169
- PMCID: PMC446105
- DOI: 10.1128/JVI.78.15.7969-7983.2004
Intracellular processing of human herpesvirus 6 glycoproteins Q1 and Q2 into tetrameric complexes expressed on the viral envelope
Abstract
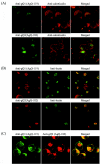
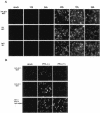
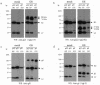
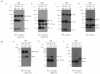
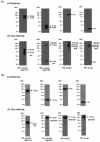
Human herpesvirus 6 (HHV-6) glycoproteins H and L (gH and gL, respectively) and the 80-kDa form of glycoprotein Q (gQ-80K) form a heterotrimeric complex that is found on the viral envelope and that is a viral ligand for human CD46. Besides gQ-80K, the gQ gene encodes an additional product whose mature molecular mass is 37 kDa (gQ-37K) and which is derived from a different transcript. Therefore, we designated gQ-80K as gQ1 and gQ-37K as gQ2. We show here that gQ2 also interacts with the gH-gL-gQ1 complex in HHV-6-infected cells and in virions. To examine how these components interact in HHV-6-infected cells, we performed pulse-chase studies. The results demonstrated that gQ2-34K, which is endo-beta-N-acetylglucosaminidase H sensitive and which is the precursor form of gQ2-37K, associates with gQ1-74K, which is the precursor form of gQ1-80K, within 30 min of the pulse period. After a 1-h chase, these precursor forms had associated with the gH-gL dimer. Interestingly, an anti-gH monoclonal antibody coimmunoprecipitated mainly gQ1-80K and gQ2-37K, with little gQ1-74K or gQ2-34K. These results indicate that although gQ2-34K and gQ1-74K interact in the endoplasmic reticulum, the gH-gL-gQ1-80K-gQ2-37K heterotetrameric complex arises in the post-endoplasmic reticulum compartment. The mature complex is subsequently incorporated into viral particles.
Figures







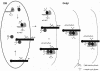
References
-
- Ablashi, D. V., N. Balachandran, S. F. Josephs, C. L. Hung, G. R. Krueger, B. Kramarsky, S. Z. Salahuddin, and R. C. Gallo. 1991. Genomic polymorphism, growth properties, and immunologic variations in human herpesvirus-6 isolates. Virology 184:545-552. - PubMed
-
- Anderson, R. A., D. X. Liu, and U. A. Gompels. 1996. Definition of a human herpesvirus-6 betaherpesvirus-specific domain in glycoprotein gH that governs interaction with glycoprotein gL: substitution of human cytomegalovirus glycoproteins permits group-specific complex formation. Virology 217:517-526. - PubMed
-
- Campadelli-Fiume, G., S. Guerrini, X. Liu, and L. Foa-Tomasi. 1993. Monoclonal antibodies to glycoprotein B differentiate human herpesvirus 6 into two clusters, variants A and B. J. Gen. Virol. 74:2257-2262. - PubMed
-
- Chandran, B., S. Tirawatnapong, B. Pfeiffer, and D. V. Ablashi. 1992. Antigenic relationships among human herpesvirus-6 isolates. J. Med. Virol. 37:247-254. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

