Calicivirus 3C-like proteinase inhibits cellular translation by cleavage of poly(A)-binding protein
- PMID: 15254188
- PMCID: PMC446144
- DOI: 10.1128/JVI.78.15.8172-8182.2004
Calicivirus 3C-like proteinase inhibits cellular translation by cleavage of poly(A)-binding protein
Abstract
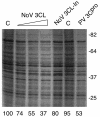
Caliciviruses are single-stranded RNA viruses that cause a wide range of diseases in both humans and animals, but little is known about the regulation of cellular translation during infection. We used two distinct calicivirus strains, MD145-12 (genus Norovirus) and feline calicivirus (FCV) (genus Vesivirus), to investigate potential strategies used by the caliciviruses to inhibit cellular translation. Recombinant 3C-like proteinases (r3CL(pro)) from norovirus and FCV were found to cleave poly(A)-binding protein (PABP) in the absence of other viral proteins. The norovirus r3CL(pro) PABP cleavage products were indistinguishable from those generated by poliovirus (PV) 3C(pro) cleavage, while the FCV r3CL(pro) products differed due to cleavage at an alternate cleavage site 24 amino acids downstream of one of the PV 3C(pro) cleavage sites. All cleavages by calicivirus or PV proteases separated the C-terminal domain of PABP that binds translation factors eIF4B and eRF3 from the N-terminal RNA-binding domain of PABP. The effect of PABP cleavage by the norovirus r3CL(pro) was analyzed in HeLa cell translation extracts, and the presence of r3CL(pro) inhibited translation of both endogenous and exogenous mRNAs. Translation inhibition was poly(A) dependent, and replenishment of the extracts with PABP restored translation. Analysis of FCV-infected feline kidney cells showed that the levels of de novo cellular protein synthesis decreased over time as virus-specific proteins accumulated, and cleavage of PABP occurred in virus-infected cells. Our data indicate that the calicivirus 3CL(pro), like PV 3C(pro), mediates the cleavage of PABP as part of its strategy to inhibit cellular translation. PABP cleavage may be a common mechanism among certain virus families to manipulate cellular translation.
Figures


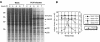


References
-
- Allaire, M., M. M. Chernaia, B. A. Malcolm, and M. N. G. James. 1994. Picornaviral 3C cysteine proteinases have a fold similar to chymotrypsin-like serine proteinases. Nature 369:72-76. - PubMed
-
- Al-Molawi, N., V. A. Beardmore, M. J. Carter, G. E. Kass, and L. O. Roberts. 2003. Caspase-mediated cleavage of the feline calicivirus capsid protein. J. Gen. Virol. 84:1237-1244. - PubMed
-
- Asselbergs, F., W. Peters, W. van Venrooij, and H. Bloemendal. 1978. Diminished sensitivity of re-initiation of translation to inhibition by cap analogues in reticulocyte lysates. Eur. J. Biochem. 88:483-488. - PubMed
-
- Belliot, G., S. V. Sosnovtsev, T. Mitra, C. Hammer, M. Garfield, and K. Y. Green. 2003. In vitro proteolytic processing of the MD145 norovirus ORF1 nonstructural polyprotein yields stable precursors and products similar to those detected in calicivirus-infected cells. J. Virol. 77:10957-10974. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

