cis-Regulatory and protein evolution in orthologous and duplicate genes
- PMID: 15256508
- PMCID: PMC509261
- DOI: 10.1101/gr.2662504
cis-Regulatory and protein evolution in orthologous and duplicate genes
Abstract
The relationship between protein and regulatory sequence evolution is a central question in molecular evolution. It is currently not known to what extent changes in gene expression are coupled with the evolution of protein coding sequences, or whether these changes differ among orthologs (species homologs) and paralogs (duplicate genes). Here, we develop a method to measure the extent of functionally relevant cis-regulatory sequence change in homologous genes, and validate it using microarray data and experimentally verified regulatory elements in different eukaryotic species. By comparing the genomes of Caenorhabditis elegans and C. briggsae, we found that protein and regulatory evolution is weakly coupled in orthologs but not paralogs, suggesting that selective pressure on gene expression and protein evolution is quite similar and persists for a significant amount of time following speciation but not gene duplication. Additionally, duplicates of both species exhibit a dramatic acceleration of both regulatory and protein evolution compared to orthologs, suggesting increased directional selection and/or relaxed selection on both gene expression patterns and protein function in duplicate genes.
Copyright 2004 Cold Spring Harbor Laboratory Press ISSN
Figures
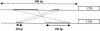
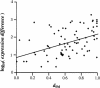
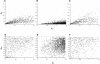
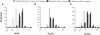
References
-
- Blencowe, B.J. 2000. Exonic splicing enhancers: Mechanism of action, diversity and role in human genetic diseases. Trends Biochem. Sci. 25: 106–110. - PubMed
-
- Castillo-Davis, C.I. and Hartl, D.L. 2002. Genome evolution and developmental constraint in Caenorhabditis elegans. Mol. Biol. Evol. 19: 728–735. - PubMed
-
- The C. elegans Sequencing Consortium. 1998. Genome sequence of the nematode C. elegans: A platform for investigating biology. Science 282: 2012–2018. - PubMed
WEB SITE REFERENCES
-
- http://wormbase.org; Wormbase.
-
- http://www.oeb.harvard.edu/faculty/wakeley/; C source code of the SMM software (sharmot).
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
