Identification of genes required for embryo development in Arabidopsis
- PMID: 15266054
- PMCID: PMC519041
- DOI: 10.1104/pp.104.045179
Identification of genes required for embryo development in Arabidopsis
Abstract
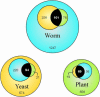
A long-term goal of Arabidopsis research is to define the minimal gene set needed to produce a viable plant with a normal phenotype under diverse conditions. This will require both forward and reverse genetics along with novel strategies to characterize multigene families and redundant biochemical pathways. Here we describe an initial dataset of 250 EMB genes required for normal embryo development in Arabidopsis. This represents the first large-scale dataset of essential genes in a flowering plant. When compared with 550 genes with other knockout phenotypes, EMB genes are enriched for basal cellular functions, deficient in transcription factors and signaling components, have fewer paralogs, and are more likely to have counterparts among essential genes of yeast (Saccharomyces cerevisiae) and worm (Caenorhabditis elegans). EMB genes also represent a valuable source of plant-specific proteins with unknown functions required for growth and development. Analyzing such unknowns is a central objective of genomics efforts worldwide. We focus here on 34 confirmed EMB genes with unknown functions, demonstrate that expression of these genes is not embryo-specific, validate a strategy for identifying interacting proteins through complementation with epitope-tagged proteins, and discuss the value of EMB genes in identifying novel proteins associated with important plant processes. Based on sequence comparison with essential genes in other model eukaryotes, we identify 244 candidate EMB genes without paralogs that represent promising targets for reverse genetics. These candidates should facilitate the recovery of additional genes required for seed development.
Figures



Similar articles
-
The importance of being essential: EMBRYO-DEFECTIVE genes in Arabidopsis.C R Biol. 2008 Oct;331(10):726-36. doi: 10.1016/j.crvi.2008.07.014. Epub 2008 Sep 2. C R Biol. 2008. PMID: 18926486 Review.
-
Genome-wide identification of EMBRYO-DEFECTIVE (EMB) genes required for growth and development in Arabidopsis.New Phytol. 2020 Apr;226(2):306-325. doi: 10.1111/nph.16071. Epub 2019 Sep 18. New Phytol. 2020. PMID: 31334862 Review.
-
Integrating the genetic and physical maps of Arabidopsis thaliana: identification of mapped alleles of cloned essential (EMB) genes.PLoS One. 2009 Oct 8;4(10):e7386. doi: 10.1371/journal.pone.0007386. PLoS One. 2009. PMID: 19812694 Free PMC article.
-
The EMB 506 gene encodes a novel ankyrin repeat containing protein that is essential for the normal development of Arabidopsis embryos.Plant J. 1999 Jan;17(2):169-79. doi: 10.1046/j.1365-313x.1999.00361.x. Plant J. 1999. PMID: 10074714
-
Arabidopsis emb175 and other ppr knockout mutants reveal essential roles for pentatricopeptide repeat (PPR) proteins in plant embryogenesis.Planta. 2005 Jun;221(3):424-36. doi: 10.1007/s00425-004-1452-x. Epub 2005 Jan 13. Planta. 2005. PMID: 15647901
Cited by
-
Transcriptome Analysis Reveals Key Seed-Development Genes in Common Buckwheat (Fagopyrum esculentum).Int J Mol Sci. 2019 Sep 3;20(17):4303. doi: 10.3390/ijms20174303. Int J Mol Sci. 2019. PMID: 31484314 Free PMC article.
-
A nuclear-encoded protein, mTERF6, mediates transcription termination of rpoA polycistron for plastid-encoded RNA polymerase-dependent chloroplast gene expression and chloroplast development.Sci Rep. 2018 Aug 9;8(1):11929. doi: 10.1038/s41598-018-30166-6. Sci Rep. 2018. PMID: 30093718 Free PMC article.
-
A Plastid-Bound Ankyrin Repeat Protein Controls Gametophyte and Early Embryo Development in Arabidopsis thaliana.Front Plant Sci. 2022 Mar 8;13:767339. doi: 10.3389/fpls.2022.767339. eCollection 2022. Front Plant Sci. 2022. PMID: 35350296 Free PMC article.
-
ATM-mediated transcriptional and developmental responses to gamma-rays in Arabidopsis.PLoS One. 2007 May 9;2(5):e430. doi: 10.1371/journal.pone.0000430. PLoS One. 2007. PMID: 17487278 Free PMC article.
-
A novel plant-specific family gene, ROOT PRIMORDIUM DEFECTIVE 1, is required for the maintenance of active cell proliferation.Plant Physiol. 2006 Feb;140(2):591-602. doi: 10.1104/pp.105.074724. Epub 2006 Jan 11. Plant Physiol. 2006. PMID: 16407439 Free PMC article.
References
-
- Albert S, Despres B, Guilleminot J, Bechtold N, Pelletier G, Delseny M, Devic M (1999) The EMB 506 gene encodes a novel ankyrin repeat containing protein that is essential for the normal development of Arabidopsis embryos. Plant J 17: 169–179 - PubMed
-
- Alonso JM, Stepanova AN, Leisse TJ, Kim CJ, Chen H, Shinn P, Stevenson DK, Zimmerman J, Barajas P, Cheuk R, et al (2003) Genome-wide insertional mutagenesis of Arabidopsis thaliana. Science 301: 653–657 - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

