Characterization of the mouse adeno-associated virus AAVS1 ortholog
- PMID: 15280500
- PMCID: PMC479059
- DOI: 10.1128/JVI.78.16.8917-8921.2004
Characterization of the mouse adeno-associated virus AAVS1 ortholog
Abstract
The nonpathogenic human adeno-associated virus (AAV) has developed a mechanism to integrate its genome into human chromosome 19 at 19q13.4 (termed AAVS1), thereby establishing latency. Here, we provide evidence that the chromosomal signals required for site-specific integration are conserved in the mouse genome proximal to the recently identified Mbs85 gene. These sequence motifs can be specifically nicked by the viral Rep protein required for the initiation of site-specific AAV DNA integration. Furthermore, these signals can serve as a minimal origin for Rep-dependent DNA replication. In addition, we isolated the mouse Mbs85 proximal promoter and show transcriptional activity in three mouse cell lines.
Figures

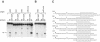
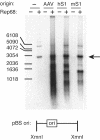
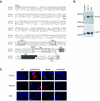
References
-
- Amiss, T. J., and R. J. Samulski. 2001. Methods for adeno-associated virus-mediated gene transfer into muscle. Methods Mol. Biol. 175:455-469. - PubMed
-
- Bakowska, J. C., M. V. Di Maria, S. M. Camp, Y. Wang, P. D. Allen, and X. O. Breakefield. 2003. Targeted transgene integration into transgenic mouse fibroblasts carrying the full-length human AAVS1 locus mediated by HSV/AAV rep(+) hybrid amplicon vector. Gene Ther. 10:1691-1702. - PubMed
-
- Berns, K. I. 1996. Parvoviridae: the viruses and their replication, p. 2173-2197. In B. N. Fields, D. M. Knipe, and P. M. Howley (ed.), Fields virology, 3rd ed., vol. 2. Lippincott-Raven, Philadelphia, Pa.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

