Transcriptionally competent chromatin assembled with exogenous histones in a yeast whole cell extract
- PMID: 15282330
- PMCID: PMC506827
- DOI: 10.1093/nar/gnh107
Transcriptionally competent chromatin assembled with exogenous histones in a yeast whole cell extract
Abstract
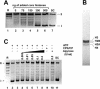
We describe a cell-free chromatin assembly system derived from the yeast Saccharomyces cerevisiae, which efficiently packages DNA into minichromosomes in a reaction dependent on exogenous core histones and an ATP-regenerating system. Both supercoiled and relaxed plasmid DNA serve as templates for nucleosomal loading in a gradual process that takes at least 6 h for completion at 30 degrees C. Micrococcal nuclease digestion of the assembled minichromosomes displays an extended nucleosomal ladder with a repeat length of 165 bp. The purified minichromosomes contain the four core histones in stoichiometric proportion and exhibit phased nucleosomes over the mouse mammary tumour virus (MMTV) promoter. The progesterone receptor and NF1 synergize on these minichromosomes resulting in efficient cell-free transcription. The ease of manipulation and the potential use of yeast strains carrying mutations in the chromatin handling machinery make this system suitable for detailed mechanistic studies.
Figures

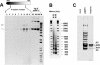

Similar articles
-
Assembly of MMTV promoter minichromosomes with positioned nucleosomes precludes NF1 access but not restriction enzyme cleavage.Nucleic Acids Res. 1998 Aug 15;26(16):3657-66. doi: 10.1093/nar/26.16.3657. Nucleic Acids Res. 1998. PMID: 9685480 Free PMC article.
-
Chromatin assembly in a crude fraction from yeast cells.Methods Mol Biol. 2006;313:209-23. doi: 10.1385/1-59259-958-3:209. Methods Mol Biol. 2006. PMID: 16118436
-
Yeast chromatin reconstitution system using purified yeast core histones and yeast nucleosome assembly protein-1.Protein Expr Purif. 1997 Jun;10(1):132-40. doi: 10.1006/prep.1996.0716. Protein Expr Purif. 1997. PMID: 9179300
-
Chromatin and transcription in yeast.Genetics. 2012 Feb;190(2):351-87. doi: 10.1534/genetics.111.132266. Genetics. 2012. PMID: 22345607 Free PMC article. Review.
-
Noncovalent modification of chromatin: different remodeled products with different ATPase domains.Cold Spring Harb Symp Quant Biol. 2004;69:183-92. doi: 10.1101/sqb.2004.69.183. Cold Spring Harb Symp Quant Biol. 2004. PMID: 16117648 Review. No abstract available.
Cited by
-
Electrophoretic Analysis of DNA Supercoiling Activities.Methods Mol Biol. 2025;2881:259-270. doi: 10.1007/978-1-0716-4280-1_13. Methods Mol Biol. 2025. PMID: 39704948
-
A method for genome-wide analysis of DNA helical tension by means of psoralen-DNA photobinding.Nucleic Acids Res. 2010 Oct;38(19):e182. doi: 10.1093/nar/gkq687. Epub 2010 Aug 4. Nucleic Acids Res. 2010. PMID: 20685815 Free PMC article.
-
Histone H1 subtypes differentially modulate chromatin condensation without preventing ATP-dependent remodeling by SWI/SNF or NURF.PLoS One. 2009 Oct 1;4(10):e0007243. doi: 10.1371/journal.pone.0007243. PLoS One. 2009. PMID: 19794910 Free PMC article.
-
Topoisomerase-modulated genome-wide DNA supercoiling domains colocalize with nuclear compartments and regulate human gene expression.Nat Struct Mol Biol. 2025 Jan;32(1):48-61. doi: 10.1038/s41594-024-01377-5. Epub 2024 Aug 16. Nat Struct Mol Biol. 2025. PMID: 39152238
References
-
- Felsenfeld G. (1992) Chromatin as an essential part of the transcriptional mechanism. Nature, 355, 219–224. - PubMed
-
- Kornberg R.D. and Lorch,Y. (1995) Interplay between chromatin structure and transcription. Curr. Opin. Cell. Biol., 7, 371–375. - PubMed
-
- Cairns B.R. (1998) Chromatin remodelling machines: similar motors, ulterior motives. Trends Biochem. Sci., 23, 20–25. - PubMed
-
- Kingston R.E. and Narlikar,G.J. (1999) ATP-dependent remodelling and acetylation as regulators of chromatin fluidity. Genes Dev., 13, 2339–2352. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

