Staphylococcus aureus DNA ligase: characterization of its kinetics of catalysis and development of a high-throughput screening compatible chemiluminescent hybridization protection assay
- PMID: 15283677
- PMCID: PMC1133749
- DOI: 10.1042/BJ20040054
Staphylococcus aureus DNA ligase: characterization of its kinetics of catalysis and development of a high-throughput screening compatible chemiluminescent hybridization protection assay
Abstract
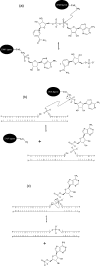
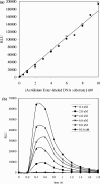
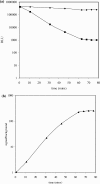
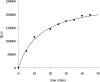
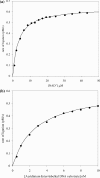
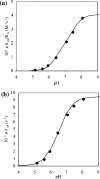
DNA ligases are key enzymes involved in the repair and replication of DNA. Prokaryotic DNA ligases uniquely use NAD+ as the adenylate donor during catalysis, whereas eukaryotic enzymes use ATP. This difference in substrate specificity makes the bacterial enzymes potential targets for therapeutic intervention. We have developed a homogeneous chemiluminescence-based hybridization protection assay for Staphylococcus aureus DNA ligase that uses novel acridinium ester technology and demonstrate that it is an alternative to the commonly used radiometric assays for ligases. The assay has been used to determine a number of kinetic constants for S. aureus DNA ligase catalysis. These included the K(m) values for NAD+ (2.75+/-0.1 microM) and the acridinium-ester-labelled DNA substrate (2.5+/-0.2 nM). A study of the pH-dependencies of kcat, K(m) and kcat/K(m) has revealed values of kinetically influential ionizations within the enzyme-substrate complexes (kcat) and free enzyme (kcat/K(m)). In each case, the curves were shown to be composed of one kinetically influential ionization, for k(cat), pK(a)=6.6+/-0.1 and kcat/K(m), pK(a)=7.1+/-0.1. Inhibition characteristics of the enzyme against two Escherichia coli DNA ligase inhibitors have also been determined with IC50 values for these being 3.30+/-0.86 microM for doxorubicin and 1.40+/-0.07 microM for chloroquine diphosphate. The assay has also been successfully miniaturized to a sufficiently low volume to allow it to be utilized in a high-throughput screen (384-well format; 20 microl reaction volume), enabling the assay to be used in screening campaigns against libraries of compounds to discover leads for further drug development.
Figures


Similar articles
-
Cloning and functional characterization of an NAD(+)-dependent DNA ligase from Staphylococcus aureus.J Bacteriol. 2001 May;183(10):3016-24. doi: 10.1128/JB.183.10.3016-3024.2001. J Bacteriol. 2001. PMID: 11325928 Free PMC article.
-
Specific and potent inhibition of NAD+-dependent DNA ligase by pyridochromanones.J Biol Chem. 2003 Oct 10;278(41):39435-42. doi: 10.1074/jbc.M306479200. Epub 2003 Jul 15. J Biol Chem. 2003. PMID: 12867414
-
Acid-base catalysis in the chemical mechanism of inosine monophosphate dehydrogenase.Biochemistry. 1999 Apr 6;38(14):4433-40. doi: 10.1021/bi9829579. Biochemistry. 1999. PMID: 10194364
-
DNA and RNA ligases: structural variations and shared mechanisms.Curr Opin Struct Biol. 2008 Feb;18(1):96-105. doi: 10.1016/j.sbi.2007.12.008. Epub 2008 Feb 8. Curr Opin Struct Biol. 2008. PMID: 18262407 Review.
-
Rapid detection of genetic mutations using the chemiluminescent hybridization protection assay (HPA): overview and comparison with other methods.Crit Rev Clin Lab Sci. 1998 Sep;35(5):369-414. doi: 10.1080/10408369891234228. Crit Rev Clin Lab Sci. 1998. PMID: 9791746 Review.
Cited by
-
Profiling the selectivity of DNA ligases in an array format with mass spectrometry.Nucleic Acids Res. 2010 Jan;38(1):e2. doi: 10.1093/nar/gkp827. Epub 2009 Oct 23. Nucleic Acids Res. 2010. PMID: 19854942 Free PMC article.
-
Mechanistic assessment of DNA ligase as an antibacterial target in Staphylococcus aureus.Antimicrob Agents Chemother. 2012 Aug;56(8):4095-102. doi: 10.1128/AAC.00215-12. Epub 2012 May 14. Antimicrob Agents Chemother. 2012. PMID: 22585221 Free PMC article.
-
Novel bacterial NAD+-dependent DNA ligase inhibitors with broad-spectrum activity and antibacterial efficacy in vivo.Antimicrob Agents Chemother. 2011 Mar;55(3):1088-96. doi: 10.1128/AAC.01181-10. Epub 2010 Dec 28. Antimicrob Agents Chemother. 2011. PMID: 21189350 Free PMC article.
References
-
- Lehman I. R. DNA ligase: structure, mechanism, and function. Science. 1974;186:790–797. - PubMed
-
- Cherepanov A. V., de Vries S. Dynamic mechanism of nick recognition by DNA ligase. Eur. J. Biochem. 2002;269:5993–5999. - PubMed
-
- Wilkinson A., Day J., Bowater R. Bacterial DNA ligases. Mol. Microbiol. 2001;40:1241–1248. - PubMed
-
- Timson D. J., Singleton M. R., Wigley D. B. DNA ligases in the repair and replication of DNA. Mutat. Res. 2000;460:301–318. - PubMed
-
- Engler M. J., Richardson C. C. The Enzymes. New York: Academic Press; 1982. pp. 3–29.
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

