Studies of SpoIIAB mutant proteins elucidate the mechanisms that regulate the developmental transcription factor sigmaF in Bacillus subtilis
- PMID: 15294015
- PMCID: PMC1134100
- DOI: 10.1042/BJ20040923
Studies of SpoIIAB mutant proteins elucidate the mechanisms that regulate the developmental transcription factor sigmaF in Bacillus subtilis
Abstract
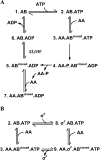
SigmaF, the first compartment-specific sigma factor of sporulation, is regulated by an anti-sigma factor, SpoIIAB (AB) and its antagonist SpoIIAA (AA). AB can bind to sigmaF in the presence of ATP or to AA in the presence of ADP; in addition, AB can phosphorylate AA. The ability of AB to switch between its two binding partners regulates sigmaF. Early in sporulation, AA activates sigmaF by releasing it from its complex with AB. We have previously proposed a reaction scheme for the phosphorylation of AA by AB which accounts for AA's regulatory role. A crucial feature of this scheme is a conformational change in AB that accompanies its switch in binding partner. In the present study, we have studied three AB mutants, all of which have amino-acid replacements in the nucleotide-binding region; AB-E104K (Glu104-->Lys) and AB-T49K (Thr49-->Lys) fail to activate sigmaF, and AB-R105A (Arg105-->Ala) activates it prematurely. We used techniques of enzymology, surface plasmon resonance and fluorescence spectroscopy to analyse the defects in each mutant. AB-E104K was deficient in binding to AA, AB-T49K was deficient in binding to ADP and AB-R105A bound ADP exceptionally strongly. Although the release of sigmaF from all three mutant proteins was impaired, and all three failed to undergo the wild-type conformational change when switching binding partners, the phenotypes of the mutant cells were best accounted for by the properties of the respective AB species in forming complexes with AA and ADP. The behaviour of the mutants enables us to propose convincing mechanisms for the regulation of sigmaF in wild-type bacteria.
Figures
Similar articles
-
Signalling network with a bistable hysteretic switch controls developmental activation of the sigma transcription factor in Bacillus subtilis.Mol Microbiol. 2006 Jul;61(1):165-84. doi: 10.1111/j.1365-2958.2006.05212.x. Mol Microbiol. 2006. PMID: 16824103
-
The kinase activity of the antisigma factor SpoIIAB is required for activation as well as inhibition of transcription factor sigmaF during sporulation in Bacillus subtilis.J Mol Biol. 1998 Dec 4;284(3):569-78. doi: 10.1006/jmbi.1998.2202. J Mol Biol. 1998. PMID: 9826499
-
Evidence for common sites of contact between the antisigma factor SpoIIAB and its partners SpoIIAA and the developmental transcription factor sigmaF in Bacillus subtilis.J Mol Biol. 1998 Dec 4;284(3):557-68. doi: 10.1006/jmbi.1998.2201. J Mol Biol. 1998. PMID: 9826498
-
Differential gene expression in genetically identical sister cells: the initiation of sporulation in Bacillus subtilis.Mol Microbiol. 2005 May;56(3):578-89. doi: 10.1111/j.1365-2958.2005.04594.x. Mol Microbiol. 2005. PMID: 15819616 Review.
-
Where asymmetry in gene expression originates.Mol Microbiol. 2005 Aug;57(3):611-20. doi: 10.1111/j.1365-2958.2005.04687.x. Mol Microbiol. 2005. PMID: 16045607 Review.
Cited by
-
Structural Analysis of the Hanks-Type Protein Kinase YabT From Bacillus subtilis Provides New Insights in its DNA-Dependent Activation.Front Microbiol. 2019 Jan 8;9:3014. doi: 10.3389/fmicb.2018.03014. eCollection 2018. Front Microbiol. 2019. PMID: 30671027 Free PMC article.
-
Differential mechanisms of binding of anti-sigma factors Escherichia coli Rsd and bacteriophage T4 AsiA to E. coli RNA polymerase lead to diverse physiological consequences.J Bacteriol. 2008 May;190(10):3434-43. doi: 10.1128/JB.01792-07. Epub 2008 Mar 21. J Bacteriol. 2008. PMID: 18359804 Free PMC article.
References
-
- Kroos L., Zhang B., Ichikawa H., Yu Y. T. Control of sigma factor activity during Bacillus subtilis sporulation. Mol. Microbiol. 1999;31:1285–1294. - PubMed
-
- Piggot P. J., Losick R. Sporulation genes and intercompartmental regulation. In: Sonenshein A. L., Hoch J. A., Losick R., editors. Bacillus subtilis and its Closest Relatives: from Genes to Cells. Washington, DC: ASM Press, American Society for Microbiology; 2002. pp. 483–517.
-
- Min K. T., Hilditch C. M., Diederich B., Errington J., Yudkin M. D. σF, the first compartment-specific transcription factor of B. subtilis, is regulated by an anti-sigma factor that is also a protein kinase. Cell. 1993;74:735–742. - PubMed
-
- Alper S., Duncan L., Losick R. An adenosine nucleotide switch controlling the activity of a cell type-specific transcription factor in B. subtilis. Cell. 1994;77:195–205. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

