Targeting the A site RNA of the Escherichia coli ribosomal 30 S subunit by 2'-O-methyl oligoribonucleotides: a quantitative equilibrium dialysis binding assay and differential effects of aminoglycoside antibiotics
- PMID: 15294017
- PMCID: PMC1134060
- DOI: 10.1042/BJ20040246
Targeting the A site RNA of the Escherichia coli ribosomal 30 S subunit by 2'-O-methyl oligoribonucleotides: a quantitative equilibrium dialysis binding assay and differential effects of aminoglycoside antibiotics
Abstract
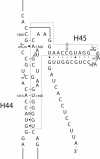
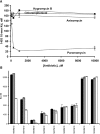
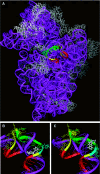
The bacterial ribosome comprises 30 S and 50 S ribonucleoprotein subunits, contains a number of binding sites for known antibiotics and is an attractive target for selection of novel antibacterial agents. On the 30 S subunit, for example, the A site (aminoacyl site) close to the 3'-end of 16 S rRNA is highly important in the decoding process. Binding by some aminoglycoside antibiotics to the A site leads to erroneous protein synthesis and is lethal for bacteria. We targeted the A site on purified 30 S ribosomal subunits from Escherichia coli with a set of overlapping, complementary OMe (2'-O-methyl) 10-mer oligoribonucleotides. An equilibrium dialysis technique was applied to measure dissociation constants of these oligonucleotides. We show that there is a single high-affinity region, spanning from A1493 to C1510 (Kd, 29-130 nM), flanked by two lower-affinity regions, within a span from U1485 to G1516 (Kd, 310-4300 nM). Unexpectedly, addition of the aminoglycoside antibiotic paromomycin (but not hygromycin B) caused a dose-dependent increase of up to 7.5-fold in the binding of the highest affinity 10-mer 1493 to 30 S subunits. Oligonucleotides containing residues complementary to A1492 and/or A1493 showed particularly marked stimulation of binding by paromomycin. The results are consistent with high-resolution structures of antibiotic binding to the A site and with greater accessibility of residues of A1492 and A1493 upon paromomycin binding. 10-mer 1493 binding is thus a probe of the conformational switch to the 'closed' conformation triggered by paromomycin that is implicated in the discrimination by 30 S subunits of cognate from non-cognate tRNA and the translational misreading caused by paromomycin. Finally, we show that OMe oligonucleotides targeted to the A site are moderately good inhibitors of in vitro translation and that there is a limited correlation of inhibition activity with binding strength to the A site.
Figures



Similar articles
-
Interaction of translation initiation factor IF1 with the E. coli ribosomal A site.J Mol Biol. 2000 May 26;299(1):1-15. doi: 10.1006/jmbi.2000.3672. J Mol Biol. 2000. PMID: 10860719
-
Effect of mutations in the A site of 16 S rRNA on aminoglycoside antibiotic-ribosome interaction.J Mol Biol. 1999 Feb 12;286(1):33-43. doi: 10.1006/jmbi.1998.2446. J Mol Biol. 1999. PMID: 9931247
-
Fluorescence-based approach for detecting and characterizing antibiotic-induced conformational changes in ribosomal RNA: comparing aminoglycoside binding to prokaryotic and eukaryotic ribosomal RNA sequences.J Am Chem Soc. 2004 Mar 24;126(11):3447-53. doi: 10.1021/ja030568i. J Am Chem Soc. 2004. PMID: 15025471
-
The A-minor motifs in the decoding recognition process.Biochimie. 2006 Aug;88(8):993-9. doi: 10.1016/j.biochi.2006.05.018. Epub 2006 Jun 19. Biochimie. 2006. PMID: 16889885 Review.
-
Improving on nature: antibiotics that target the ribosome.Curr Opin Microbiol. 2005 Oct;8(5):534-42. doi: 10.1016/j.mib.2005.08.004. Curr Opin Microbiol. 2005. PMID: 16111914 Review.
Cited by
-
Antibiotic Resistance via Bacterial Cell Shape-Shifting.mBio. 2022 Jun 28;13(3):e0065922. doi: 10.1128/mbio.00659-22. Epub 2022 May 26. mBio. 2022. PMID: 35616332 Free PMC article.
-
Antibiotic drugs targeting bacterial RNAs.Acta Pharm Sin B. 2014 Aug;4(4):258-65. doi: 10.1016/j.apsb.2014.06.012. Epub 2014 Jul 31. Acta Pharm Sin B. 2014. PMID: 26579393 Free PMC article. Review.
-
Interactions of 2'-O-methyl oligoribonucleotides with the RNA models of the 30S subunit A-site.PLoS One. 2018 Jan 19;13(1):e0191138. doi: 10.1371/journal.pone.0191138. eCollection 2018. PLoS One. 2018. PMID: 29351348 Free PMC article.
-
Crystal structures of complexes between aminoglycosides and decoding A site oligonucleotides: role of the number of rings and positive charges in the specific binding leading to miscoding.Nucleic Acids Res. 2005 Oct 7;33(17):5677-90. doi: 10.1093/nar/gki862. Print 2005. Nucleic Acids Res. 2005. PMID: 16214802 Free PMC article.
-
Thermal Stability of Peptide Nucleic Acid Complexes.J Phys Chem B. 2019 Oct 3;123(39):8168-8177. doi: 10.1021/acs.jpcb.9b05168. Epub 2019 Sep 20. J Phys Chem B. 2019. PMID: 31491077 Free PMC article.
References
-
- Ramakrishnan V. Ribosome structure and the mechanism of translation. Cell. 2002;108:557–572. - PubMed
-
- Spahn C. M. T., Prescott C. D. Throwing a spanner in the works: antibiotics and the translational apparatus. J. Mol. Med. 1996;74:423–439. - PubMed
-
- Neu H. C., Gootz T. D. Antimicrobial chemotherapy. In: Baron S., editor. Medical Microbiology, chapter 11. Galveston: University of Texas Medical Branch; 1996. - PubMed
-
- Carter A. P., Clemons W. M., Brodersen D. E., Morgan-Warren R. J., Wimberley B. T., Ramakrishnan V. Functional insights from the structure of the 30 S ribosomal subunit and its interactions with antibiotics. Nature (London) 2000;407:340–348. - PubMed
-
- Odom O. W., Picking W. D., Tsalkova T., Hardesty B. The synthesis of polyphenylalanine on ribosomes to which erythromycin is bound. Eur. J. Biochem. 1991;198:713–722. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

