Guidelines for genotyping in genomewide linkage studies: single-nucleotide-polymorphism maps versus microsatellite maps
- PMID: 15311375
- PMCID: PMC1182056
- DOI: 10.1086/424696
Guidelines for genotyping in genomewide linkage studies: single-nucleotide-polymorphism maps versus microsatellite maps
Abstract
Genomewide linkage scans have traditionally employed panels of microsatellite markers spaced at intervals of approximately 10 cM across the genome. However, there is a growing realization that a map of closely spaced single-nucleotide polymorphisms (SNPs) may offer equal or superior power to detect linkage, compared with low-density microsatellite maps. We performed a series of simulations to calculate the information content associated with microsatellite and SNP maps across a range of different marker densities and heterozygosities for sib pairs (with and without parental genotypes), sib trios, and sib quads. In the case of microsatellite markers, we varied density across 11 levels (1 marker every 0.5, 1, 2, 3, 4, 5, 6, 7, 8, 9, or 10 cM) and marker heterozygosity across 6 levels (2, 3, 4, 5, 10, or 20 equally frequent alleles), whereas, in the case of SNPs, we varied marker density across 4 levels (1 marker every 0.1, 0.2, 0.5, or 1 cM) and minor-allele frequency across 7 levels (0.5, 0.4, 0.3, 0.2, 0.1, 0.05, and 0.01). When parental genotypes were available, a map consisting of microsatellites spaced every 2 cM or a relatively sparse map of SNPs (i.e., at least 1 SNP/cM) was sufficient to extract most of the inheritance information from the map (>95% in most cases). However, when parental genotypes were unavailable, it was important to use as dense a map of markers as possible to extract the greatest amount of inheritance information. It is important to note that the information content associated with a traditional map of microsatellite markers (i.e., 1 marker every ~10 cM) was significantly lower than the information content associated with a dense map of SNPs or microsatellites. These results strongly suggest that previous linkage studies that employed sparse microsatellite maps could benefit substantially from reanalysis by use of a denser map of markers.
Figures
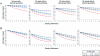

References
-
- Abecasis GR, Cherny SS, Cardon LR (2001) The impact of genotyping error on family-based analysis of quantitative traits. Eur J Hum Genet 9:130–134 - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

