OmpT: molecular dynamics simulations of an outer membrane enzyme
- PMID: 15315948
- PMCID: PMC1304768
- DOI: 10.1529/biophysj.104.046987
OmpT: molecular dynamics simulations of an outer membrane enzyme
Abstract
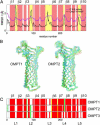
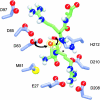
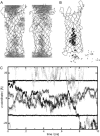
Five molecular dynamics simulations (total duration >25 ns) have been performed on the Escherichia coli outer membrane protease OmpT embedded in a dimyristoylphosphatidylcholine lipid bilayer. Globally the protein is conformationally stable. Some degree of tilt of the beta-barrel is observed relative to the bilayer plane. The greatest degree of conformational flexibility is seen in the extracellular loops. A complex network of fluctuating H-bonds is formed between the active site residues, such that the Asp210-His212 interaction is maintained throughout, whereas His212 and Asp83 are often bridged by a water molecule. This supports a catalytic mechanism whereby Asp83 and His212 bind a water molecule that attacks the peptide carbonyl. A configuration yielded by docking calculations of OmpT simulation snapshots and a model substrate peptide Ala-Arg-Arg-Ala was used as the starting point for an extended Huckel calculation on the docked peptide. These placed the lowest unoccupied molecular orbital mainly on the carbon atom of the central C=O in the scissile peptide bond, thus favoring attack on the central peptide by the water held by residues Asp83 and His212. The trajectories of water molecules reveal exchange of waters between the intracellular face of the membrane and the interior of the barrel but no exchange at the extracellular mouth. This suggests that the pore-like region in the center of OmpT may enable access of water to the active site from below. The simulations appear to reveal the presence of specific lipid interaction sites on the surface of the OmpT barrel. This reveals the ability of extended MD simulations to provide meaningful information on protein-lipid interactions.
Figures



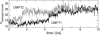
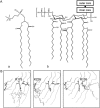
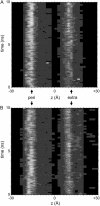
References
-
- Achouak, W., T. Heulin, and J. M. Pages. 2001. Multiple facets of bacterial porins. FEMS Microbiol. Lett. 199:1–7. - PubMed
-
- Arora, A., F. Abildgaard, J. H. Bushweller, and L. K. Tamm. 2001. Structure of outer membrane protein A transmembrane domain by NMR spectroscopy. Nat. Struct. Biol. 8:334–338. - PubMed
-
- Baaden, M., C. Meier, and M. S. P. Sansom. 2003. A molecular dynamics investigation of mono- and dimeric states of the outer membrane enzyme OMPLA. J. Mol. Biol. 331:177–189. - PubMed
-
- Basyn, F., B. Charloteaux, A. Thomas, and R. Brasseur. 2001. Prediction of membrane protein orientation in lipid bilayers: a theoretical approach. J. Mol. Graphics Mod. 20:235–244. - PubMed
-
- Berendsen, H. J. C., J. P. M. Postma, W. F. van Gunsteren, A. DiNola, and J. R. Haak. 1984. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81:3684–3690.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

