Membrane position of a basic aromatic peptide that sequesters phosphatidylinositol 4,5 bisphosphate determined by site-directed spin labeling and high-resolution NMR
- PMID: 15315949
- PMCID: PMC1304792
- DOI: 10.1529/biophysj.104.046748
Membrane position of a basic aromatic peptide that sequesters phosphatidylinositol 4,5 bisphosphate determined by site-directed spin labeling and high-resolution NMR
Abstract
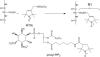
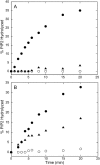
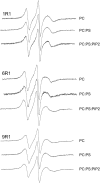
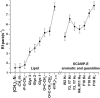
The membrane interactions and position of a positively charged and highly aromatic peptide derived from a secretory carrier membrane protein (SCAMP) are examined using magnetic resonance spectroscopy and several biochemical methods. This peptide (SCAMP-E) is shown to bind to membranes containing phosphatidylinositol 4,5-bisphosphate, PI(4,5)P2, and sequester PI(4,5)P2 within the plane of the membrane. Site-directed spin labeling of the SCAMP-E peptide indicates that the position and structure of membrane bound SCAMP-E are not altered by the presence of PI(4,5)P2, and that the peptide backbone is positioned within the lipid interface below the level of the lipid phosphates. A second approach using high-resolution NMR was used to generate a model for SCAMP-E bound to bicelles. This approach combined oxygen enhancements of nuclear relaxation with a computational method to dock the SCAMP-E peptide at the lipid interface. The model for SCAMP generated by NMR is consistent with the results of site-directed spin labeling and places the peptide backbone in the bilayer interfacial region and the aromatic side chains within the lipid hydrocarbon region. The charged side chains of SCAMP-E lie well within the interface with two arginine residues lying deeper than a plane defined by the position of the lipid phosphates. These data suggest that SCAMP-E interacts with PI(4,5)P2 through an electrostatic mechanism that does not involve specific lipid-peptide contacts. This interaction may be facilitated by the position of the positively charged side chains on SCAMP-E within a low-dielectric region of the bilayer interface.
Figures




Similar articles
-
Secretory carrier membrane protein SCAMP2 and phosphatidylinositol 4,5-bisphosphate interactions in the regulation of dense core vesicle exocytosis.Biochemistry. 2007 Sep 25;46(38):10909-20. doi: 10.1021/bi701121j. Epub 2007 Aug 22. Biochemistry. 2007. PMID: 17713930
-
Location of the myristoylated alanine-rich C-kinase substrate (MARCKS) effector domain in negatively charged phospholipid bicelles.Biophys J. 2003 Oct;85(4):2442-8. doi: 10.1016/S0006-3495(03)74667-0. Biophys J. 2003. PMID: 14507707 Free PMC article.
-
Membrane-bound orientation and position of the synaptotagmin I C2A domain by site-directed spin labeling.Biochemistry. 2003 Jan 14;42(1):96-105. doi: 10.1021/bi0268145. Biochemistry. 2003. PMID: 12515543
-
The measurement of immersion depth and topology of membrane proteins by solution state NMR.Biochim Biophys Acta. 2007 Dec;1768(12):3044-51. doi: 10.1016/j.bbamem.2007.09.011. Epub 2007 Oct 16. Biochim Biophys Acta. 2007. PMID: 17976526 Review.
-
Experimental nuclear magnetic resonance studies of membrane proteins.Methods Enzymol. 1994;239:536-60. doi: 10.1016/s0076-6879(94)39021-5. Methods Enzymol. 1994. PMID: 7830598 Review. No abstract available.
Cited by
-
SCAMP2 interacts with Arf6 and phospholipase D1 and links their function to exocytotic fusion pore formation in PC12 cells.Mol Biol Cell. 2005 Oct;16(10):4463-72. doi: 10.1091/mbc.e05-03-0231. Epub 2005 Jul 19. Mol Biol Cell. 2005. PMID: 16030257 Free PMC article.
-
Solid-state NMR (31)P paramagnetic relaxation enhancement membrane protein immersion depth measurements.J Phys Chem B. 2014 Apr 24;118(16):4370-7. doi: 10.1021/jp500267y. Epub 2014 Apr 11. J Phys Chem B. 2014. PMID: 24689497 Free PMC article.
-
The role of hydrophobic interactions in positioning of peripheral proteins in membranes.BMC Struct Biol. 2007 Jun 29;7:44. doi: 10.1186/1472-6807-7-44. BMC Struct Biol. 2007. PMID: 17603894 Free PMC article.
-
Secretory carrier membrane proteins.Protoplasma. 2012 Apr;249(2):269-83. doi: 10.1007/s00709-011-0295-0. Epub 2011 Jun 2. Protoplasma. 2012. PMID: 21633931 Review.
-
SCAMP3 negatively regulates epidermal growth factor receptor degradation and promotes receptor recycling.Mol Biol Cell. 2009 Mar;20(6):1816-32. doi: 10.1091/mbc.e08-09-0894. Epub 2009 Jan 21. Mol Biol Cell. 2009. PMID: 19158374 Free PMC article.
References
-
- Altenbach, C., K. J. Oh, R. J. Trabanino, K. Hideg, and W. L. Hubbell. 2001. Estimation of inter-residue distances in spin labeled proteins at physiological temperatures: experimental strategies and practical limitations. Biochemistry. 40:15471–15482. - PubMed
-
- Arbuzova, A., L. Wang, J. Wang, G. Hangyas-Mihalyne, D. Murray, B. Honig, and S. McLaughlin. 2000. Membrane binding of peptides containing both basic and aromatic residues. Experimental studies with peptides corresponding to the scaffolding region of caveolin and the effector region of MARCKS. Biochemistry. 39:10330–10339. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

